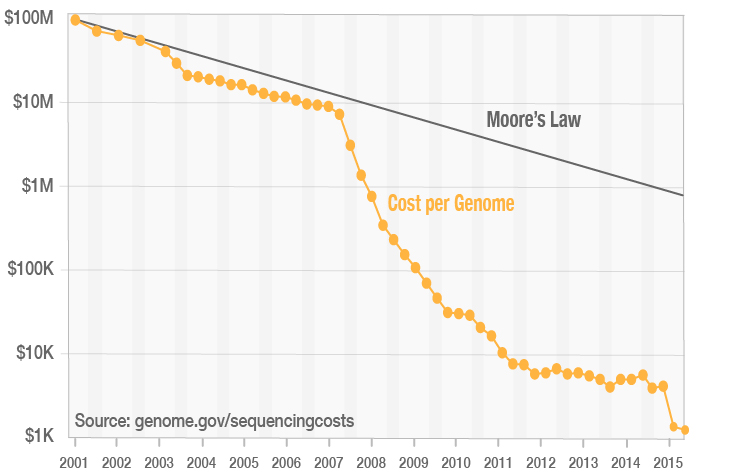
Cost of Next-Generation Sequencing
Redefining the Price of Discovery
Since the completion of the Human Genome Project, the cost of next-generation sequencing (NGS) has decreased at a dramatic rate, outpacing Moore’s Law. Through continuous innovation, Illumina has helped reduce the cost of NGS, enabling the $1000 human genome.
As next-generation sequencing costs continue to decline, Illumina is leading the way in making NGS more affordable and accessible. We strive to help labs of all sizes access the potential of this powerful technology. With these resources, we’ll guide you through key factors to consider when planning your NGS budget.
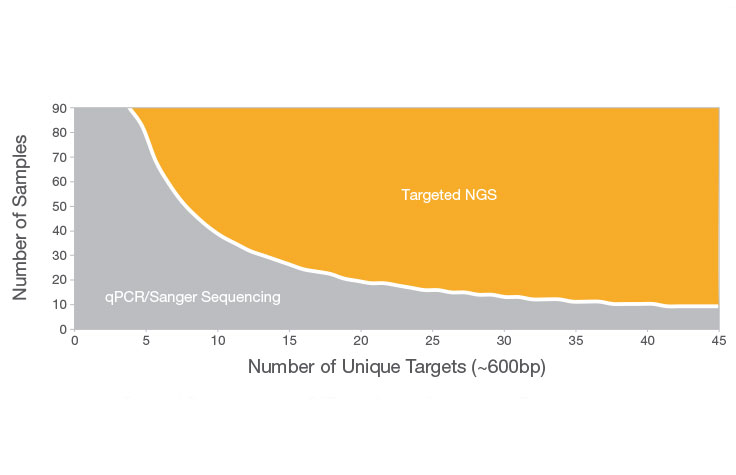
Next-Generation Sequencing Cost Comparison
When evaluating NGS costs, consider the sample volume for your study. In general, for analyzing only a few (< 20) targets on a few samples, traditional methods such as Sanger sequencing or qPCR can be useful. For sequencing more than 20 target regions or high sample volumes, NGS is preferable.
NGS also delivers higher discovery power and sensitivity to detect novel or rare variants. It offers a hypothesis-free approach that doesn’t require prior knowledge of sequence information. These insights can be invaluable for enabling discovery and fueling research publications.
Learn more about:
NGS vs. qPCR
NGS vs. Sanger sequencing
NGS vs. microarrays
Next-Generation Sequencing Cost Considerations
When estimating the cost of NGS, consider these factors:
- Instrument purchase
- Cost per sample (eg, DNA isolation, library prep, and sequencing reagents)
- Data analysis and storage
- Optional instrument support plan
- Optional training and preventive maintenance
Also consider additional lab equipment, such as:
- Nucleic acid quantitation instrument
- Nucleic acid quality analyzer
- Thermocycler
- Ultrasonicator (used in some library prep methods)
- Centrifuge
- Common lab supplies (eg, pipettors, 96-well plates, centrifuge tubes)
Buyer's Guide to NGS Systems
Find tips to help you estimate next-generation sequencing costs and choose the right instrument for your lab.
Download GuideNGS Cost Consultation
Have questions about how next-generation sequencing fits into your budget? Connect with an Illumina representative.
Contact Us
Examples of NGS Cost Per Sample
| Application | Estimated Cost Per Sample | Experimental Parameters |
|---|---|---|
| Targeted gene expression profiling | $23 USD | Cost per sample calculation is based on a run using:
|
| 16S metagenomic sequencing | $18 USD | Cost per sample calculation is based on a run using:
|
Low-Cost NGS Instrument
Designed for simplicity, the iSeq 100 Sequencing System makes next-generation sequencing easier and more affordable than ever. Discover more—without the cost.
View System
Cost of NGS Data Analysis
The cost of NGS data storage and analysis is one of the most common questions for beginners. Three factors will influence your data analysis budget:
- Licensing a data analysis software platform
- Storing data (often part of the licensing cost)
- Running data analysis apps (also known as the compute cost)
The compute cost can vary depending on the amount of sequencing data you analyze. To help you calculate the cost of NGS data analysis for your study, we've estimated the volume of data generated for common methods.
| Application | Estimated Data Output |
|---|---|
| Human whole-genome sequencing (at 30× coverage) | ~120 Gb |
| Human exome sequencing (at 100× coverage) | ~8 Gb |
| Microbial whole-genome sequencing | ~300 Mb |
| 16S rRNA sequencing | ~60 Mb |
1 megabase (Mb) = 1,000,000 bases
1 gigabase (Gb) = 1,000,000,000 bases
The volume of data generated is related to the sequencing coverage level for your experiment.
Learn More
Free Trial for NGS Data Analysis
Try data analysis apps in BaseSpace Sequence Hub free for 30 days, without instrument purchase. You’ll also receive 250 complimentary iCredits to cover additional storage or compute costs.
Learn More About iCreditsNGS Experimental Design
Learn about read length, sequencing coverage, and more—everything you need for your first sequencing run.
Start Planning