Introduction
There is significant variation between individuals in the rate of drug metabolism for a large number of clinically prescribed drugs. A strong contributing factor to this variability in drug metabolism is the genetic composition of the drug-metabolizing enzymes, and thus genotyping the genes that encode these enzymes (i.e. pharmacogenes) is an important component of personalized medicine1. Cytochrome P450 2D6 (CYP2D6) encodes one of the most important drug-metabolizing enzymes and is responsible for the metabolism of about 21% of clinically used drugs2. The CYP2D6 gene is highly polymorphic, with more than 100 star alleles defined by the Pharmacogene Variation (PharmVar) Consortium3. Star alleles4 are gene haplotypes defined by a combination of small variants (SNVs and indels) and structural variants (SVs), and correspond to different levels of enzymatic activity, ie, poor, intermediate, normal, or ultrarapid metabolizer5–7.
Genotyping CYP2D6 is challenged by common deletions and duplications of CYP2D6 and hybrids between CYP2D6 and its pseudogene paralog, CYP2D74,8,9, which shares 94% sequence similarity, including a few near-identical regions8,10. Traditionally, CYP2D6 genotyping is done in low or medium throughput with array-based platforms, such as the PharmacoScan, or polymerase chain reaction (PCR) based methods such as TaqMan assays, ddPCR and long-range PCR. These assays differ in the number of star alleles (variants) they interrogate, leading to variability in genotyping results across assays8,11,12. To detect SVs, these assays or test platforms may need to be complemented with Copy Number Variation (CNV) assays that may also be limited to detection of just a subset of the known CNVs4,9.
With recent advances in next-generation sequencing, it is now possible to profile the entire genome at high-throughput and in a clinically-relevant timeframe. Driven by these advances, many countries are undertaking large scale population sequencing projects13–15 wherein pharmacogenomics (PGx) testing will greatly increase the clinical utility of these efforts. To enable PGx within these studies, we developed a CYP2D6 caller, Cyrius16, a novel WGS-specific CYP2D6 genotyping tool that overcomes the challenges with the homology between CYP2D6 and CYP2D7. Here we provide an overview of Cyrius and future directions for applications of PGx enabled by WGS.
CYP2D6 genotyping method used by Cyrius
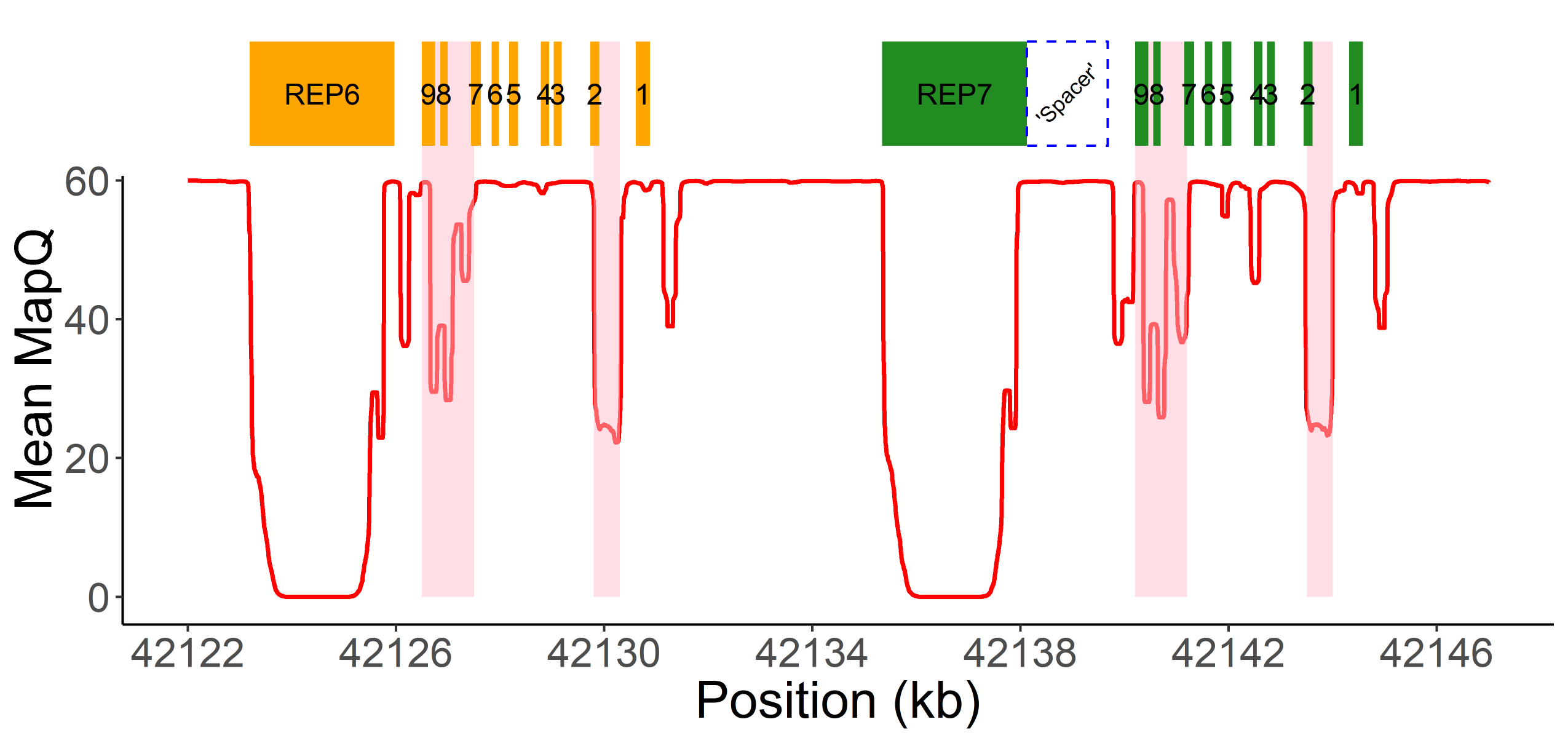
Read alignment accuracy is reduced in CYP2D6 because of the homology with CYP2D7 (Figure 1) and this can make variant calling challenging and error prone. Cyrius uses a novel approach to overcome this challenge and a detailed workflow is illustrated in Figure 2 using the Coriell sample NA12878 (*3/*68+*4) as an example. First, Cyrius identifies the total copies of both CYP2D6 and CYP2D7 combined by calculating the read depth from the WGS aligned BAM file using all reads mapped to either CYP2D6 or CYP2D7. Next Cyrius determines the number of complete CYP2D6 genes as well as identify hybrid genes based on 117 bases that have been demonstrated to differ between CYP2D6 and CYP2D7 in the population. Hybrids are identified when the copy number (CN) of CYP2D6 measured at these bases changes within the gene. For example, the sample NA12878 shown in Figure 2B has two full copies of CYP2D6 and one hybrid where Exon 1 comes from CYP2D6 and Exons 2-9 come from CYP2D7 (ie, *68). Next Cyrius parses the read alignments to identify the protein-changing small variants that define star alleles (Figure 2C). Finally, Cyrius matches the SVs and small variants against star allele definitions and produces genotypes that are consistent with the called variants.
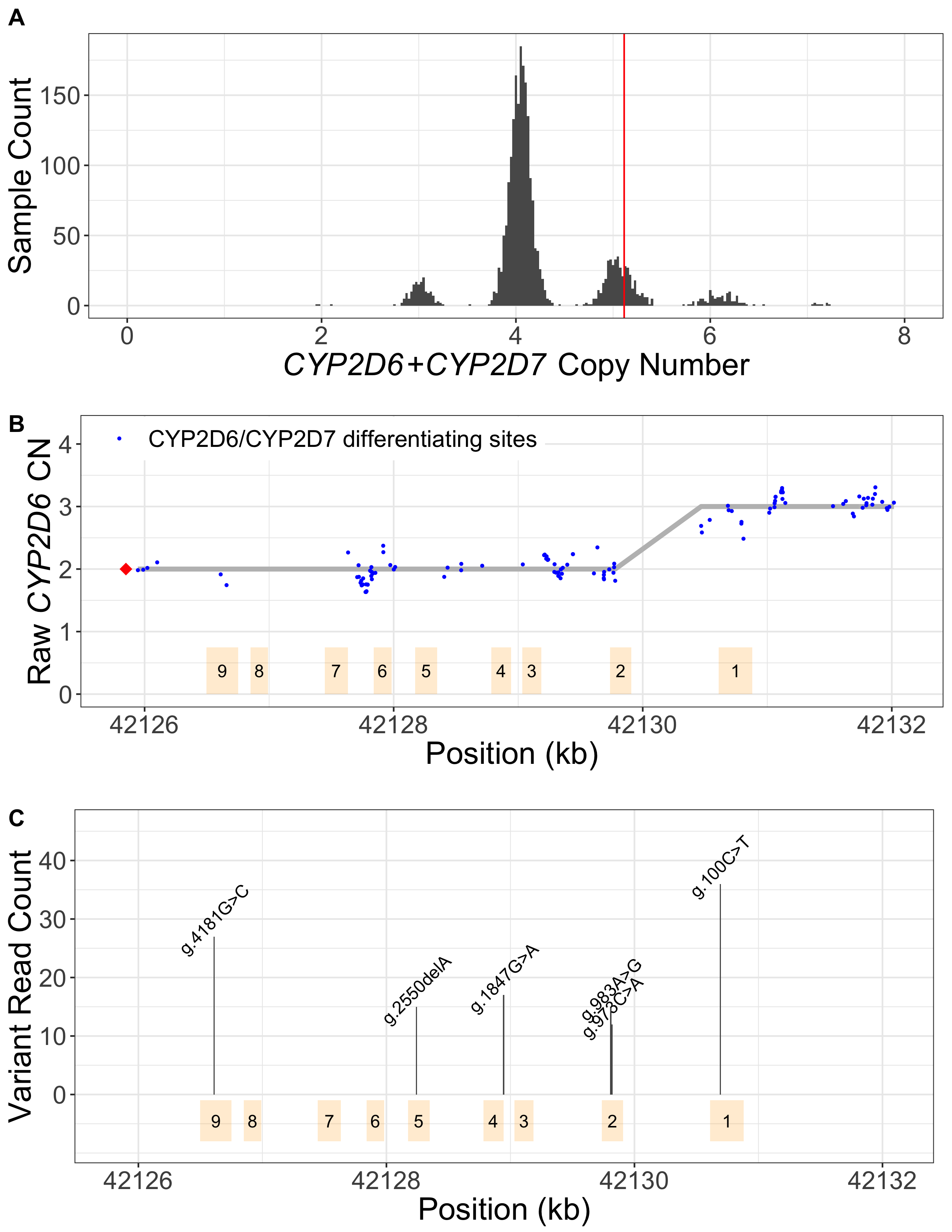
A. CN(CYP2D6+CYP2D7) is derived by counting and modelling all reads that align to either CYP2D6 or CYP2D7. The histogram shows the distribution of normalized CYP2D6+CYP2D7 depth in 1kGP samples, showing peaks at CN2, 3, 4, 5, 6 and 7. The red vertical line represents the value for NA12878, corresponding to CN5 that indicates an additional copy (could be CYP2D6 or hybrid). B. SVs are called by examining the CNs of the CYP2D6/CYP2D7 differentiating bases. Exons are denoted by yellow boxes. Blue dots denote raw CYP2D6 CNs, calculated as CN(CYP2D6+CYP2D7) multiplied by the ratio of CYP2D6 supporting reads out of CYP2D6 and CYP2D7 supporting reads. The red diamond denotes the CN of genes that are CYP2D6-derived at the 3’ end (can be complete CYP2D6 or CYP2D7-CYP2D6 hybrid), calculated as CN(CYP2D6+CYP2D7) minus CN(spacer). The CYP2D6 CN is called at each CYP2D6/CYP2D7 differentiating site and a change in CYP2D6 CN within the gene indicates the presence of a hybrid. In NA12878, the CYP2D6 CN changes from 2 to 3 between Exon 2 and Exon 1, indicating a CYP2D6-CYP2D7 hybrid (*68). C. Supporting read counts of the star-allele defining protein-changing small variants are used to call the CN of each variant. The y axis shows the read counts for all queried small variant positions. Six variants are called in NA12878, one of which, g.100C>T, is called as two copies (one copy belongs to *4 and the other belongs to *68 based on star allele definitions). Finally, star alleles are called based on detected SVs and small variants.
Performance validation
The availability of a panel of reference samples by the CDC Genetic Testing Reference Material Program (GeT-RM)11,17, where the consensus genotypes of major pharmacogenes are derived using multiple genotyping platforms, has enabled assessment of genotyping accuracy for newly developed methods. We assessed the CYP2D6 calls made by Cyrius against 144 truthset samples, including 138 GeT-RM samples and 8 samples with truth generated using PacBio HiFi sequence reads (two samples overlap between GeT-RM and PacBio). Cyrius initially made 5 discordant calls in the 144 truth samples, showing a concordance of 96.5%. We were subsequently able to identify the causes and improve Cyrius to correctly call 4 of these 5 samples, reaching a ‘trained’ concordance of 99.3% (143 out of 144 samples). Cyrius’s accuracy is higher than two existing CYP2D6 callers Aldy18 (86.8%) and Stargazer19 (84.0%)16.
Together, the validation samples used in this study confirmed our CYP2D6 calling accuracy in 47 distinct haplotypes, including 40 star alleles as well as several SV structures, such as duplications and tandem arrangements including *13+*2, *68+*4, *36+*10 and *36+*36+*10. These 40 star alleles represent 30.5% of the 131 star alleles in PharmVar (as of July 2020) and 51.7% (31 out of 60) of the star alleles with known function. Applying Cyrius to the samples from the 1000 Genomes Project (1kGP)20, representing five distinct populations (Europeans, Africans, East Asians, South Asians and admixed Americans), indicates that these 40 validated star alleles represent 96% of the star alleles present in the pangenomic population.
We next assessed Mendelian consistency of the Cyrius calls in sequencing data from 597 1kGP trios. While the comparison above against truth genotypes allows for different haplotype phasing, the Mendelian consistency check is a more stringent check of the phasing of the star alleles when more than two copies of CYP2D6 are present. Of the 572 trios with calls in all three family members, 561 (98.1%) are Mendelian consistent. All of the inconsistent trios could be resolved by changing the phasing - i.e. no proband had a called star allele that was absent in both parents. The majority (8/11) of the inconsistent cases are where the trio identified that two identical copies of CYP2D6 should be on the same haplotype with the other haplotype having no copies of CYP2D6 (ie, *Cyrius call *1/*1 vs. trio-based phasing *5/*1x2). This Mendelian consistency check confirms the consistency of the genotypes across the pedigree but not the accuracy of the star alleles called. Combining the trio concordance tests with the accuracy tests performed above against truth genotypes provides confidence in the overall accuracy of the genotypes produced by Cyrius.
Actionable pharmacogenetic variants across five ethnic populations
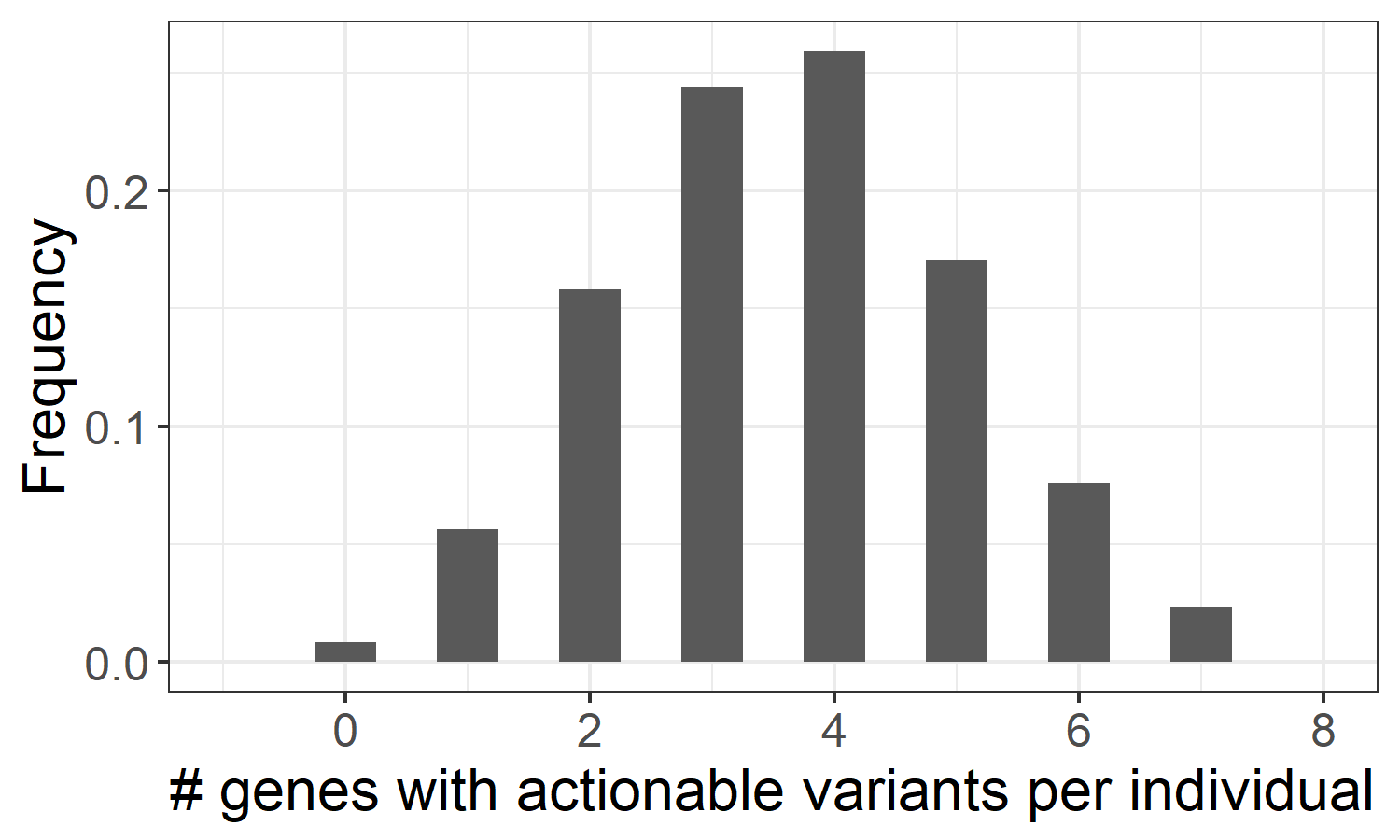
The method employed in Cyrius can be applied to resolve other paralogs that suffer from the same homology problem, including another pharmacogene CYP2B6. In order to provide a more complete pharmacogenomics picture, we have developed an additional pipeline to genotype 19 pharmacogenes with CPIC21 level A guidelines (https://cpicpgx.org/genes-drugs/), where genetic information should be used to change the prescription of affected drugs. These 19 genes include CACNA1S, CFTR, CYP2B6, CYP2C19, CYP2C9, CYP2D6, CYP3A5, CYP4F2, DPYD, G6PD, IFNL3, NUDT15, RYR1, SLCO1B1, TPMT, UGT1A1, VKORC1, HLA-A and HLA-B, among which we use specialized callers to genotype CYP2D6/CYP2B6 (Cyrius), and HLA-A/HLA-B (DRAGEN HLA caller). Translating small variants and CNVs into star alleles and activity scores allowed us to assess the distribution of the different metabolizer statuses in the 1kGP samples, and identify those genes with a genotype that requires modified drug prescription. Based on this analysis, we estimate that 99.2% of individuals in the pangenomic population have at least one pharmacogene with actionable variants (Figure 3), indicating that WGS could provide valuable pharmacogenetic information for almost every healthy individual.
Summary
We present a new software tool, Cyrius, that accurately genotypes the highly complex CYP2D6 region. Across the 144 validation samples, we are able to confirm the accuracy of Cyrius across 40 different star alleles that represent roughly 96% of the star alleles in the pangenomic population. Readers interested in learning more about Cyrius are encouraged to read our manuscript that describes this method16. Compared with traditional genotyping assays, WGS provides a promising option for building up more accurate allele frequency databases across populations because it assays all of the variants including CNVs and, combined with the right software, is able to resolve all of the known star alleles accurately. With the continued development of WGS-based PGx tools and more targeted methods like Cyrius, we can help accelerate pharmacogenomics and move one step closer towards personalized medicine.
For more information or a DRAGEN trial license for academic use, please contact dragen-info@illumina.com.
Acknowledgements
We thank the CDC Genetic Testing Reference Material Program (GeT-RM) for generating the consensus genotypes. We thank the New York Genome Center and the Coriell Institute for Medical Research for generating and releasing the 1kGP WGS data. We thank our coauthors Fei Shen, Nina Gonzaludo, Alka Malhotra, Cande Rogert, Ryan Taft and David Bentley. We thank Bochao Zhang for providing the DRAGEN HLA caller.
External links
Publication: https://www.nature.com/articles/s41397-020-00205-5
Software: https://github.com/Illumina/Cyrius
Related content
References
- Evans WE, Relling MV. Moving towards individualized medicine with pharmacogenomics. Nature.2004;429(6990):464-468. doi:10.1038/nature02626
- Zhou S-F. Polymorphism of human cytochrome P450 2D6 and its clinical significance: Part I. Clin Pharmacokinet.2009;48(11):689-723. doi:10.2165/11318030-000000000-00000
- Gaedigk A, Ingelman-Sundberg M, Miller NA, et al. The Pharmacogene Variation (PharmVar) Consortium: Incorporation of the Human Cytochrome P450 (CYP) Allele Nomenclature Database. Clin Pharmacol Ther.2018;103(3):399-401. doi:10.1002/cpt.910
- Nofziger C, Turner AJ, Sangkuhl K, et al. PharmVar GeneFocus: CYP2D6. Clin Pharmacol Ther.2020;107(1):154-170. doi:10.1002/cpt.1643
- Gaedigk A, Simon SD, Pearce RE, Bradford LD, Kennedy MJ, Leeder JS. The CYP2D6 activity score: translating genotype information into a qualitative measure of phenotype. Clin Pharmacol Ther.2008;83(2):234-242. doi:10.1038/sj.clpt.6100406
- Gaedigk A, Sangkuhl K, Whirl-Carrillo M, Klein T, Leeder JS. Prediction of CYP2D6 phenotype from genotype across world populations. Genet Med.2017;19(1):69-76. doi:10.1038/gim.2016.80
- Caudle KE, Sangkuhl K, Whirl-Carrillo M, et al. Standardizing CYP2D6 Genotype to Phenotype Translation: Consensus Recommendations from the Clinical Pharmacogenetics Implementation Consortium and Dutch Pharmacogenetics Working Group. Clin Transl Sci.2020;13(1):116-124. doi:10.1111/cts.12692
- Nofziger C, Paulmichl M. Accurately genotyping CYP2D6: not for the faint of heart. Pharmacogenomics.2018;19(13):999-1002. doi:10.2217/pgs-2018-0105
- Yang Y, Botton MR, Scott ER, Scott SA. Sequencing the CYP2D6 gene: from variant allele discovery to clinical pharmacogenetic testing. Pharmacogenomics.2017;18(7):673-685. doi:10.2217/pgs-2017-0033
- Gaedigk A. Complexities of CYP2D6 gene analysis and interpretation. Int Rev Psychiatry Abingdon Engl.2013;25(5):534-553. doi:10.3109/09540261.2013.825581
- Pratt VM, Everts RE, Aggarwal P, et al. Characterization of 137 Genomic DNA Reference Materials for 28 Pharmacogenetic Genes: A GeT-RM Collaborative Project. J Mol Diagn JMD.2016;18(1):109-123. doi:10.1016/j.jmoldx.2015.08.005
- Bousman CA, Jaksa P, Pantelis C. Systematic evaluation of commercial pharmacogenetic testing in psychiatry: a focus on CYP2D6 and CYP2C19 allele coverage and results reporting. Pharmacogenet Genomics.2017;27(11):387-393. doi:10.1097/FPC.0000000000000303
- Ashley EA. The Precision Medicine Initiative: A New National Effort. JAMA.2015;313(21):2119-2120. doi:10.1001/jama.2015.3595
- The Genome of the Netherlands Consortium, Francioli LC, Menelaou A, et al. Whole-genome sequence variation, population structure and demographic history of the Dutch population. Nat Genet.2014;46(8):818-825. doi:10.1038/ng.3021
- Turnbull C, Scott RH, Thomas E, et al. The 100 000 Genomes Project: bringing whole genome sequencing to the NHS. BMJ. 2018;361:k1687. doi:10.1136/bmj.k1687
- Chen X, Shen F, Gonzaludo N, et al. Cyrius: accurate CYP2D6 genotyping using whole-genome sequencing data. Pharmacogenomics J.Published online January 18, 2021:1-11. doi:10.1038/s41397-020-00205-5
- Gaedigk A, Turner A, Everts RE, et al. Characterization of Reference Materials for Genetic Testing of CYP2D6 Alleles: A GeT-RM Collaborative Project. J Mol Diagn JMD.Published online August 9, 2019. doi:10.1016/j.jmoldx.2019.06.007
- Numanagić I, Malikić S, Ford M, et al. Allelic decomposition and exact genotyping of highly polymorphic and structurally variant genes. Nat Commun.2018;9(1):1-11. doi:10.1038/s41467-018-03273-1
- Lee S, Wheeler MM, Patterson K, et al. Stargazer: a software tool for calling star alleles from next-generation sequencing data using CYP2D6 as a model. Genet Med.2019;21(2):361. doi:10.1038/s41436-018-0054-0
- The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature.2015;526(7571):68-74. doi:10.1038/nature15393
- Relling MV, Klein TE. CPIC: Clinical Pharmacogenetics Implementation Consortium of the Pharmacogenomics Research Network. Clin Pharmacol Ther.2011;89(3):464-467. doi:10.1038/clpt.2010.279