Public Health Surveillance
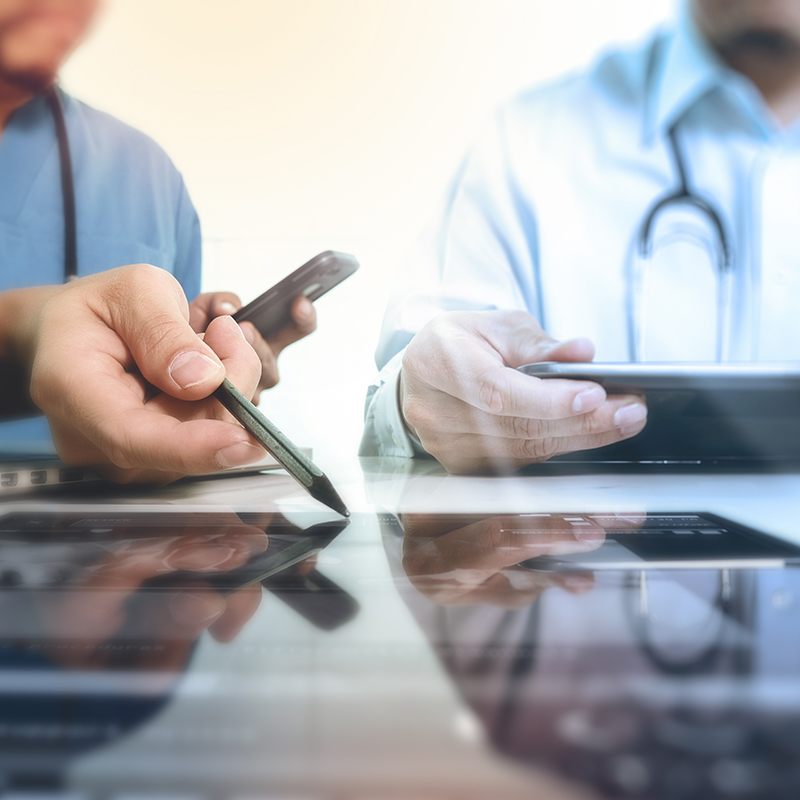

What is public health surveillance?
Public health surveillance is a vital tool for understanding the health of a population and informing public health decision-making.
It involves the systematic collection and analysis of health-related data, which can be used to identify trends and patterns, assess the effectiveness of interventions, and plan and evaluate public health programs.
A central goal of public health is to achieve equity within our communities. By concentrating on community-wide prevention and overall health, public health improves our quality of life, saves money for our communities, helps children thrive, and reduces human suffering.

Transforming public health with the power of genomic surveillance
Infectious diseases are a growing threat throughout the world. Learn how NGS is changing the way we monitor, prevent, and mitigate infectious diseases to improve surveillance and preparedness.
Download eBook
How does genomics contribute to public health surveillance?
Genomics can advance public health and wellness by detecting and characterizing new, emerging, and circulating pathogens. By tracking and monitoring infectious agent transmission and disease spread within a community, we can better understand pathogen behavior (infectivity and pathogenicity) and map the evolution of variants. This will ultimately aid the development of diagnostics and design of both therapeutic vaccines and treatments.
Advancing public health surveillance with NGS
The discriminatory power of next-generation sequencing (NGS) is advancing public health surveillance with greater speed, accuracy, and confidence than ever before. Learn more about topics within public health efforts along with diverse solutions offered from Illumina.
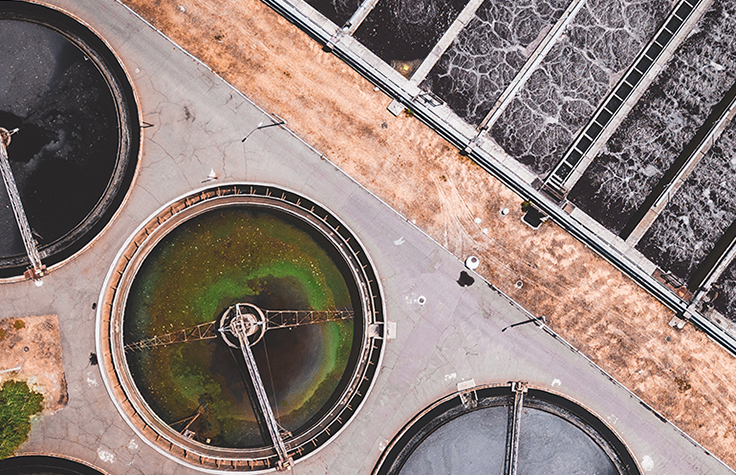

Wastewater surveillance
Wastewater surveillance allows researchers to monitor outbreaks and other threats by detecting, identifying, and characterizing pathogens in wastewater. This approach can yield important insights such as the spread of diseases for a wide variety of pathogens within a community.

Tuberculosis surveillance
Tuberculosis (TB) surveillance is an important step in addressing the second leading cause of death from infectious disease. Learn how genomics supports public health efforts against tuberculosis through detection, characterization, tracking, and monitoring.
Coronavirus surveillance
Learn how NGS can identify novel coronavirus mutations, track transmission, study viral genetics, and more with high sensitivity.
Healthcare-associated infection surveillance
Healthcare-associated infections (HAIs) are infections that occur in a healthcare setting and are often caused by multidrug-resistant organisms. These infections can lead to significant morbidity and mortality, increased healthcare costs, and regulatory noncompliance.
Antimicrobial resistance detection
Microorganisms can adapt to become resistant to antimicrobial compounds through genomic changes. Learn more about how NGS can detect low-frequency variants and genomic arrangements with unprecedented high-throughput capabilities.

NGS methods for food safety surveillance
Explore how NGS can offer substantial improvements in food safety with sensitive detection and high-throughput capabilities in areas of early detection, outbreak responses, and clinical research.
Pandemic preparedness through the power of genomic surveillance
Explore how NGS-based public health surveillance can be used to support three key pillars of pandemic preparedness for infectious diseases: early detection, outbreak response, and clinical research.
Early detection
Genomic insights using NGS can provide early detection of infectious diseases by:
- Identifying new and emerging pathogens
- Detecting pathogens in populations
- Monitoring potential reservoirs of pathogens in an environment
Outbreak responses
Infectious disease outbreak responses involve an extensive network of public health professionals. Genomic data from NGS can be used to rapidly characterize pathogens for:
- Tracking transmission information at local and national levels
- Monitoring new variants of concern
- Mapping disease clusters to inform public health priorities and interventions
Clinical research
Clinical research plays a central role in identifying and characterizing infectious diseases through the:
- Description of molecular mechanisms of infectious diseases
- Design of diagnostics and therapeutic interventions
- Development of effective vaccines

Genomic surveillance methods to aid the next pandemic prevention
Up to 75% of new or emerging infectious diseases are estimated to have zoonotic origins.1,2 We now know that zoonotic reservoirs are significant in the spread of pathogens. With NGS, it is possible to screen reservoir animals such as bats to predict and prevent viral pathogen outbreaks.
Infectious disease surveillance with NGS-methods can help us understand interspecies transmission, how zoonotic diseases emerge, spread and become resistant to common therapies, and enable us to better contain, treat and prevent disease outbreaks.
Hybrid capture sequencing can be scaled up to monitor zoonotic pathogens, while genomic surveillance with metagenomics allows for unbiased, culture-free detection and identification of a broad number of pathogens. Metagenomics also helps us understand the relationship between our microbiome and pathogen interactions, which is important when designing measures to control infectious diseases.
Learn more about:
Genomic surveillance methods
Amplicon-based library prep solutions for genomic surveillance and research
See how two new library prep solutions are enabling public health professionals to keep us safe using the proven chemistry from the COVIDSeq workflow. Watch the video to learn how the Illumina Microbial Amplicon Library Prep and the llumina Microbial Amplicon Prep—Influenza A/B are helping build, expand, and improve global genomic surveillance efforts.
NGS Workflow Finder
Take the guesswork out of your next workflow. The NGS Workflow Finder provides personalized solution recommendations and resources so you can sequence with confidence.
Find your NGS workflow todayRelated Content
Automation solutions for food safety surveillance with the MiSeq i100 Series
In this technical note, you’ll learn how an automated NGS workflow can streamline food safety surveillance and reduce hands‑on time in the lab.
Food safety surveillance with the MiSeq i100 Series
This app note highlights a complete NGS workflow for public health surveillance of foodborne pathogens. It demonstrates how Illumina DNA Prep, the MiSeq i100 Series, and DRAGEN analysis enable accurate detection of multidrug-resistant Salmonella strains.
Wastewater surveillance using the Urinary Pathogen ID/AMR Panel
This app note describes how the MiSeq i100 Series and Urinary Pathogen ID/AMR Panel offer a quick, comprehensive workflow for detecting and characterizing antibiotic resistant genes and bacterial pathogens from samples, aiding wastewater surveillance for public health.
Track emerging infectious diseases with wastewater surveillance
This application note outlines an NGS workflow for broad viral detection in wastewater surveillance, supporting public health efforts. The MiSeq i100 Series provides same-day results for a rapid response.
Can genomic surveillance detect the next pandemic before it’s too late?
Watch Dr. James Wood and Dr. William Haseltine discuss the current state of infectious disease surveillance, its strengths, and shortcomings.
Illumina sequencing of clinical samples for virus detection in a public health laboratory
Read how NGS-based methods are revolutionizing high-throughput sequencing for surveillance.
Developing a national genomic surveillance strategy
Read this guide from the World Health Organization (WHO) to understand important considerations for developing a national genomic surveillance strategy or action plan for pathogens with pandemic and epidemic potential.
Precision metagenomics eBook
Download this eBook to learn how precision metagenomics is providing innovative solutions to rapidly detect and characterize pathogens through the power of next-generation sequencing.
Related Videos
Interested in learning more about public health surveillance methods?
Fill out the form below, and we’ll be in touch.References
- Woolhouse ME, Gowtage-Sequeria S. Host range and emerging and reemerging pathogens. Emerg Infect Dis. 2005;11:1842–7. 10.3201/eid1112.050997
- Jones KE, Patel NG, Levy MA, Storeygard A, Balk D, Gittleman JL, et al. Global trends in emerging infectious diseases. Nature. 2008;451:990–3.

