Introduction
Illumina next-generation sequencing (NGS) technology uses clonal amplification and sequencing by synthesis (SBS) chemistry to enable high-throughput DNA sequencing. Used by researchers and clinicians around the globe, NGS can provide valuable information in response to numerous biological questions. There are applications in cell and molecular biology, complex disease genomics, microbiology and infectious disease, reproductive health, and more. Technological advancements have reduced the cost of NGS from ~$10M per human genome in 2007 to less than $1K today. Illumina works continuously to innovate and lower the cost of NGS while increasing throughput and accuracy to support our mission of improving human health by unlocking the power of the genome.
Illumina NGS technology is a proprietary method for DNA sequencing. Prior to sequencing, DNA fragments are captured on a lawn of oligos, bound to the surface of a flow cell and clonally amplified into clusters. Each cluster in this dense array is sequenced by incorporating fluorescently labeled deoxyribonucleotide triphosphates (dNTPs) onto the DNA strands over sequential cycles of DNA synthesis. During each cycle, the flow cell is fluorescently imaged and the emission from each cluster is recorded, allowing the nucleotides to be identified across millions of DNA fragments in a massively parallel fashion.
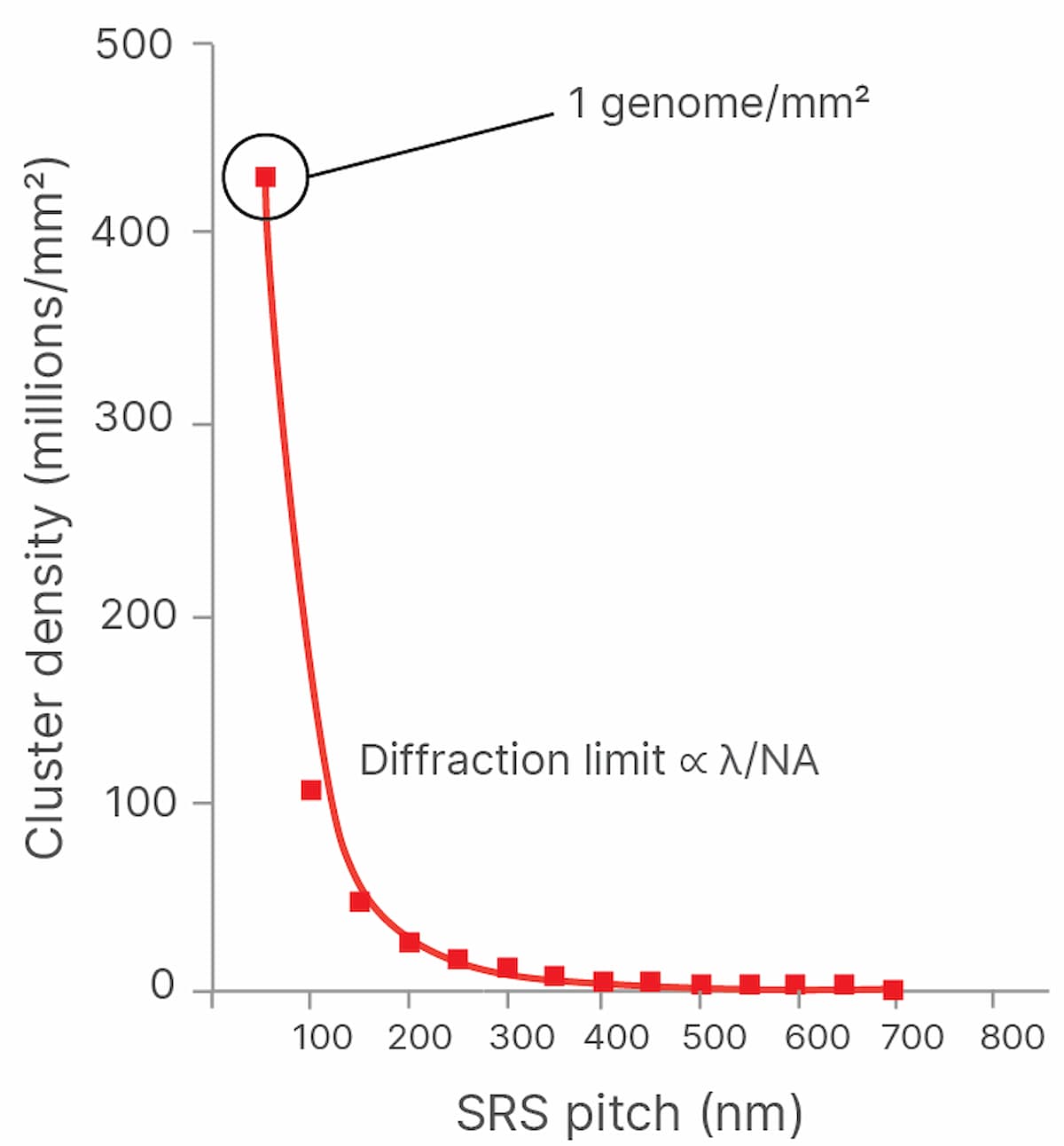
Both the speed of imaging and the density at which clusters can be packed onto a flow cell determine the throughput and cost of sequencing. The more densely the DNA clusters can be packed onto the flow cell, the more cost-effective sequencing becomes because the expensive chemical reagents flowed across the clusters produce more sequencing output per flow cell. However, DNA clusters cannot be packed arbitrarily closely together and still be imaged, as the wave nature of light and the properties of imaging systems result in a spatial resolution limit, also known as the diffraction limit, designated as λ (Figure 1).
Super-resolution optics
Illumina scientists and engineers are working to optimize the packing density of sequencing clusters. In academia, the past decade has seen a renaissance in fluorescent microscopy via the advent of new “super-resolution” imaging technologies. These technologies use various techniques to break the diffraction limit, allowing fluorescence imaging systems to reach resolutions surpassing current diffraction limits by 10-fold, reaching tens of nanometers of resolution and lower. Applying these techniques to imaging tightly packed DNA clusters on flow cells may result in lower sequencing costs (Figure 2). Super-resolution imaging techniques can be broken down into three main classes: localization-based techniques, confocal scanning imaging-based approaches, and structured illumination microscopy.
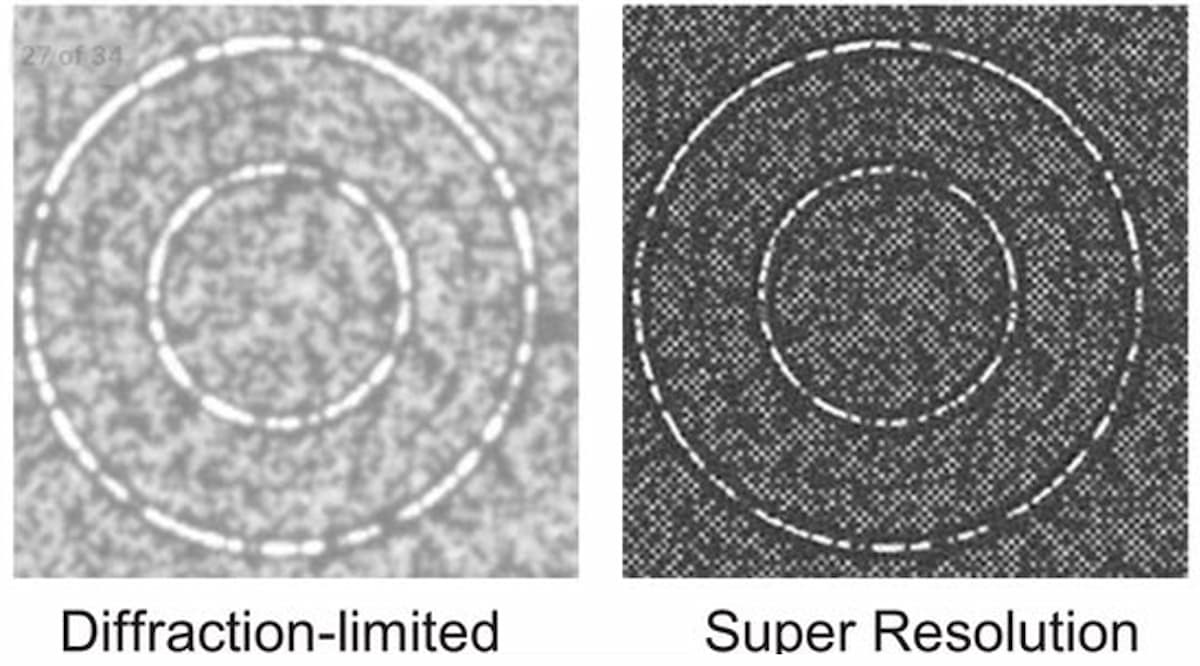

Localization-based techniques
Localization-based techniques, such as direct stochastic optical reconstruction microscopy (dSTORM), function by switching off most fluorescent molecules in a sample via strong laser excitation, which allows for the isolation of individual fluorescent molecules. These fluorescing molecules are observed as individual blurred spots on the image (point-spread functions (PSF)). The location of each molecule can be determined to tens of nanometer precision by localizing the center position of this PSF. Unfortunately, this technique requires stochastic switching of the fluorescent molecule set on and off many times so that different molecules can be localized in each frame, building up a point-like image over a long duration (> 10 mins). While this method results in very high-resolution imaging, it is slow. In addition, the need to image single molecules requires a highly sensitive imaging system and is generally limited to small fields of view, minimizing sequencing throughput. Ultimately, localization-based techniques are difficult to combine with sequencing-based imaging.
Confocal scanning imaging-based approaches
Another class of super-resolution techniques, including stimulated emission depletion (STED) microscopy, are based on the concept of confocal scanning imaging. A focused laser beam is raster scanned across a sample to build an image. The resolution of the resulting image is diffraction limited, determined by the focused spot-size (a few hundred nanometers) and the size of a filtering pinhole placed in the emission pathway. The STED concept enhances resolution via a doughnut-shaped infrared beam, which selectively deactivates fluorophores in the periphery of the focused spot via stimulated emission. This minimizes the effective area of spot illumination, enhancing the achievable resolution for the imaging system. This approach shares limitations with dSTORM: the raster-scanning aspect limits throughput, while the powerful and expensive lasers are detrimental to sample health and sequencing system economy. Thus, STED is also not ideal for use in NGS.
Structured illumination microscopy
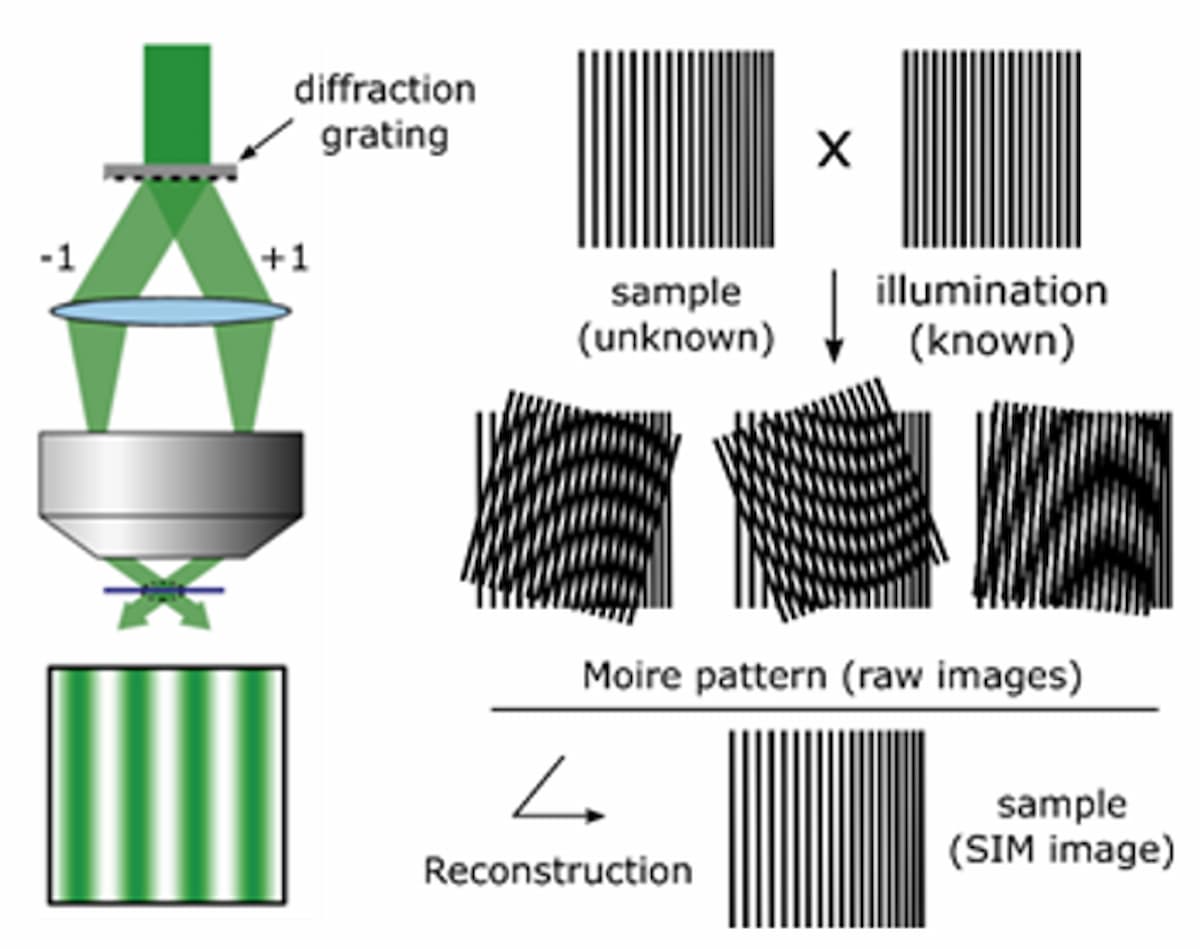
A third technique, structured illumination microscopy (SIM), works by projecting an interference pattern onto the sample to produce Moiré patterns that downshift unresolved spatial frequencies to lower, resolvable frequencies. The projected fringe pattern is usually scanned in three phases, repeated at three defined angles, to enable isotropic resolution enhancement. The resulting nine images undergo complex image processing to reconstruct the many Moiré patterns into an image with up to 2× increased resolution (Figure 3). Given that the previously mentioned techniques have distinct challenges, the unique trade-off of cost, speed, SNR, and resolution makes SIM more practical for combining with high-throughput NGS.
Speed considerations for SIM
When evaluating SIM for integration into NGS optical systems, the question of exposure time is an important consideration. Speed is a critical parameter for NGS, so the need to collect nine image exposures for a typical SIM implementation, plus the additional time to actuate the motors to shift the phase and angles of the fringes is undesirable. Furthermore, NGS imaging requires a certain signal-to-noise ratio (SNR) to produce high-quality sequencing data. How does SIM affect this? For example, in regular sequencing, a particular laser power might require an exposure time (t) to reach sufficient laser-dose and SNR. Specifically, does SIM then require a total exposure time of 9 × t? Or can the total photon budget be split between individual images of short duration, eg, t / 9 per phase image? Given the complex image processing involved in SIM, the answer is not immediately apparent. The solution requires an analysis of the way spatial frequencies and their corresponding signal-to-noise ratios are defined through the optical-transfer-function of the optics, and then manipulated and propagated within the linear algebra of the SIM algorithm, as well as the artifacts that are introduced in the process.
During investigations to optimize speed, Illumina scientists and engineers determined that the need for isotropic resolution enhancement is not a strict requirement for Illumina NGS systems. As our nanowells are arrayed in well-defined hexagonal or square patterns, the full three angles described in SIM are not a strict requirement.
Hardware implementation of SIM
The practical implementation of SIM into NGS systems can be carried out in several ways, including:
- Using a lens system to project the image of a diffraction grating or spatial light modulator onto the sample
- Splitting a laser beam into two beams and directing them onto the sample at different angles to create an interference pattern
Given that reducing the cost of sequencing is the primary goal, integrating a SIM architecture into an NGS system must be done without significant increases in hardware costs. Some academic implementations of SIM perform the rotation and phase-shifting of the fringe pattern using a spatial light modulator (SLM), which is quick and mechanically simple, but SLMs are expensive and have restrictive fields of view. Similarly, using a motor to rotate a mounted diffraction grating at sufficient speed, and with sufficient specifications, to prevent tilting of the fringes in the sample would also be cost-prohibitive. To overcome this challenge, Illumina scientists and engineers developed a rotating in-line grating system (RIGS) to produce a low-cost way of rapidly switching between angles and phases with good repeatability and rotational specification.
The RIGS system
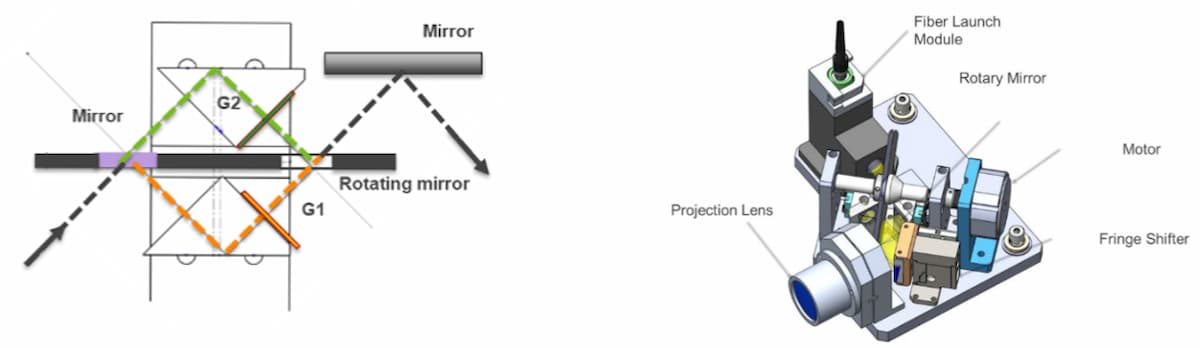
The RIGS system employs two mirrors mounted on a single rotating spindle to switch a single light source between two optical paths, each path containing gratings with the required SIM angles (Figure 4). The simple two-state system allows rapid switching between the two fringe angles without complex closed-loop control systems or calibrations. The simplicity enables repeated and reliable operation, critical to a field instrument. Furthermore, decoupling the actuation from the physical grating permits repeatable and static grating parameters (such as rotation and tilt) and robustness to thermal effects. A fast actuator downstream of the gratings is then responsible for rapid phases changes, after angle selection. In summary, the SIM imaging system can operate faster, more reliably, and with less reliance on fine-tuning of the positioning compared to systems that implement moveable gratings or other components.
Meeting stability requirements for SIM
Imaging at the nanometer scale requires comparable stability. To achieve this, many academic labs perform super-resolution imaging in stable environments (eg, the ground floor of a building) on massive optical tables equipped with costly pneumatic isolation and vibration damping. As mentioned, NGS systems need to remain cost-effective with the flexibility to be placed in regular lab environments, without the need for costly changes in lab infrastructure. Illumina scientists and engineers developed a sophisticated staging system to control rapid repositioning of optical components relative to the sample to image entire flow cells, while simultaneously maintaining precise, static positioning and vibration isolation during each imaging step, together with rapid image refocusing between motions. The required level of stability was achieved without the need for costly hardware or infrastructure, resulting in an instrument robust enough to be deployed on sea vessels.
Computational challenges of SIM
Beyond the hardware and the basic image collection process, there are further significant challenges to the economical implementation of SIM technology. SIM requires sophisticated reconstruction algorithms to transform the multiple diffraction-limited images into a single, high-resolution image. Without clever combination of algorithms and hardware, this process can be cost-prohibitive.
Resolving millions of data sources spread across an image plane presents a significant computational challenge and is often done using specialized hardware. To minimize this burden, many innovations on both the algorithm and the computational sides had to be invented. For example, Illumina scientists applied a mathematical approach to reduce the “total flops” of computation required by SIM more than eight times (2^3).
Summary
At Illumina, we are working continually to innovate and improve NGS technology to increase performance and lower costs. One area of focus is to increase the density of clusters on flow cells. However, this presents the challenge of spatially resolving clusters packed so closely together that they are below an optical system’s diffraction limit. In this article we have presented several advances employed and developed by Illumina scientists to apply super-resolution optics to NGS systems.