Wastewater Surveillance
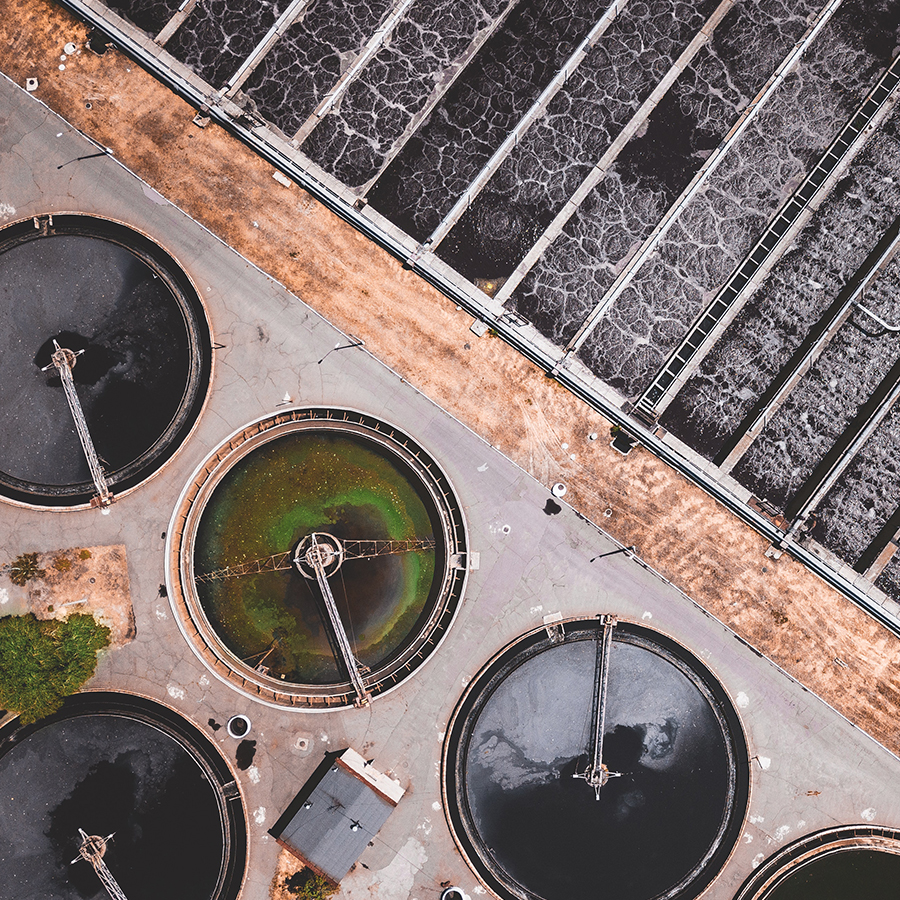

What is wastewater surveillance?
Wastewater surveillance is a method to detect, identify and characterize pathogens found in wastewater.
This method provides data to help monitor outbreaks and other threats at the community level. By knowing where these threats are, communities can better allocate resources during a public health response. Genomic information from wastewater can be used to not only determine which pathogens are present in a community, but also identify what strains and how many variants may be in the community.

How does wastewater surveillance work?
Wastewater surveillance is a process of detecting pathogens from sewage plants. Samples are collected before waste treatment and analyzed in laboratories to identify the type of pathogen and their levels in a community.1 Next-generation sequencing (NGS) can be used to analyze samples, which provides public health officials with a faster and more comprehensive way to track infections at scale. NGS can also provide information on the geographic spread of diseases, the effectiveness of community-level responses, and more.
Related App Notes
Track emerging infectious diseases with wastewater surveillance
This application note outlines an NGS workflow for broad viral detection in wastewater surveillance, supporting public health efforts.
Wastewater surveillance using the Urinary Pathogen ID/AMR Panel
This application note describes how the MiSeq i100 Series and Urinary Pathogen ID/AMR Panel offer an NGS workflow for detecting and characterizing antibiotic resistant genes (ARGs) and bacterial pathogens directly from samples to support wastewater surveillance for public health.
Precision metagenomics for rapid identification and characterization of pathogens
Download this eBook to learn how precision metagenomics is providing innovative solutions to rapidly detect and characterize pathogens through the power of next-generation sequencing.
Download eBookPCR vs NGS-based wastewater surveillance
While wastewater surveillance has been used to detect pharmaceutical, chemical, and industrial waste, it is now more important than ever to monitor wastewater for antibiotic resistance markers, bacteria, and viruses that have a significant impact on local communities. To detect pathogens of high risk to public health, RT-PCR and NGS are at the forefront of surveillance techniques.2,3 See the comparison below to learn more about each approach to wastewater sequencing.
| RT-PCR | NGS | |
| Target | Only targets small regions of the genome | Targets and characterizes whole genomes |
| Cost | Lower cost | Higher cost |
| Throughput capability | Low-throughput | High-throughput |
| Variant detection | Unable to detect variants | Can detect and identify variants |
| Data | Data is easier to interpret | Data is more complex and comprehensive |
What are some methods for wastewater surveillance?
Wastewater samples usually contain multiple pathogens and microbes,4 as opposed to clinical samples which usually only contain a single pathogen. To analyze pathogens of interest, it is recommended to use amplicon- or enrichment-based NGS library preparation methods. Amplicon-based approaches use primers to specifically target regions of microbes and/or whole genomes of single viruses, while enrichment approaches use probe hybridization to capture genomes of interest. In addition, shotgun metagenomic sequencing provides a comprehensive method to analyze all genes and microbes within a sample.5
Amplicon
Amplicon-based approaches use primers to specifically target regions of microbes and/or whole genomes of single viruses. Read how scientists are using amplicon-based methods for wastewater surveillance.
Early detection through wastewater surveillance
Read how wastewater genomic epidemiology can provide an unbiased representation of community-wide infection dynamics and surveillance to help guide public health action and policy decisions.
Identifying SARS-CoV-2 variants in wastewater
Scientists describe how targeted amplicon sequencing of wastewater samples is a powerful surveillance tool and may contribute to the early detection of emerging SARS-CoV-2 variants.
Hybrid capture enrichment
Hybridization-based enrichment is a useful strategy for analyzing specific genetic variants in a given sample, delivering dependable results across a wide range of input types and quantities. Learn more about how researchers are using hybrid capture enrichment methods for wastewater surveillance.
Monitoring antimicrobial resistance
See how wastewater-based epidemiology can complement more traditional surveillance efforts by providing community-level data to identify current and emerging antimicrobial resistance threats.
Detecting region-specific SARS-CoV-2 variants
Investigators detail how epidemiological surveillance through wastewater sequencing can aid in tracking exact viral strains in an epidemic context.
Shotgun metagenomics
Shotgun metagenomic sequencing allows researchers to comprehensively sample all genes in all organisms present in a given complex sample. Shotgun metagenomics also provides a means to study unculturable microorganisms that are otherwise difficult or impossible to analyze. See papers below to understand how shotgun metagenomics is being used for wastewater surveillance.
Using wastewater RNA-Seq to detect emerging and endemic viruses
Scientists use RNA-Seq to simultaneously detect multiple infectious diseases, providing geographical and epidemiological data to help direct healthcare interventions in India.
Metagenomic and resistome analysis
Read how scientists used metagenomic shotgun sequencing and real-time PCR to characterize the composition of bacteria and phage diversity and the resistome of major treatment processes within a wastewater treatment plant.
NGS Workflow Finder
Take the guesswork out of your next workflow. The NGS Workflow Finder provides personalized solution recommendations and resources so you can sequence with confidence.
Find your NGS workflow todayRelated Products
Viral Surveillance Panel
Obtain whole-genome sequencing (WGS) data that can characterize 66 viruses that are of high risk to public health, including SARS-CoV-2, Influenza, Mpox Virus, and Poliovirus.
Illumina COVIDSeq Test
A high-throughput, next-generation sequencing test to detect SARS-CoV-2 in patients suspected of COVID-19 and enables virus genome analysis in research use.
Respiratory Pathogen ID/AMR Enrichment Panel Kit
A next-generation sequencing (NGS)-based respiratory pathogen panel that delivers highly sensitive, comprehensive pathogen detection and antimicrobial resistance (AMR) insights.
Urinary Pathogen ID/AMR Enrichment Kit
A next-generation sequencing (NGS) panel that detects and quantifies over 170 common, less common, challenging-to-grow, and frequently missed uropathogens.
Respiratory Virus Oligo Panel
A highly sensitive NGS panel to detect and characterize common respiratory viruses, including COVID-19 strains, with comprehensive, rapid target enrichment sequencing.
Ribo-Zero Plus Microbiome rRNA Depletion Kit
A streamlined RNA-to-analysis solution for metatranscriptome sequencing of complex microbial samples, including stool.
Interested in receiving next-generation sequencing solutions for wastewater surveillance?
Fill out the form below, and we’ll be in touch.Additional Resources
Wastewater-based epidemiology
Discover the how surveillance of infectious disease through wastewater sequencing can detect SARS-CoV-2 variants and other respiratory viruses in the community in this app note.
Can wastewater surveillance protect public health?
Read the article to learn how Biobot Analytics is using wastewater data to predict COVID surges and other outbreaks.
Wastewater sequencing reveals early cryptic SARS-CoV-2 variant transmission
Learn how wasterwater surveillance is aiding in revealing early COVID transmission in this scientific publication.
Poliovirus environmental surveillance
In this publication, scientists discuss how they established an environmental surveillance program to detect non-polio enteroviruses and poliovirus in Haiti.
Using wastewater surveillance to detect COVID-19
Learn how health officials can track the spread of the virus and identify new variants, like the B.1.17 strain that became widespread using wastewater sequencing to test samples.
Related Videos
References
- Wastewater Monitoring—How Does It Work? cdc.gov/nwss/pdf/Wastewater-COVID-infographic-h.pdf. Accessed December 7, 2023.
- Science & Tech Spotlight: Wastewater Surveillance. gao.gov/products/gao-22-105841. Accessed December 7, 2023.
- Diamond MB, Keshaviah A, Bento AI, et al. Wastewater surveillance of pathogens can inform public health responses. Nature Medicine 2022; 28(10):1992-1995.
- Levy JI, Andersen KG, Knight R, et al. Wastewater surveillance for public health. Science 2023 Jan 6;379(6627):26-27.
- Wensel CR, Pluznick JL, Salzberg SL, et al. Next-generation sequencing: insights to advance clinical investigations of the microbiome. J Clin Invest. 2022 Apr 1; 132(7): e154944.