Sequencing Workflow Accuracy
Accuracy Across the Illumina Sequencing Workflow
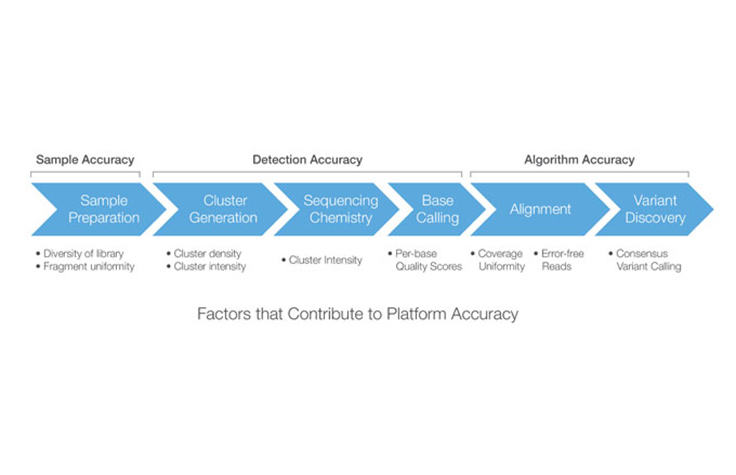
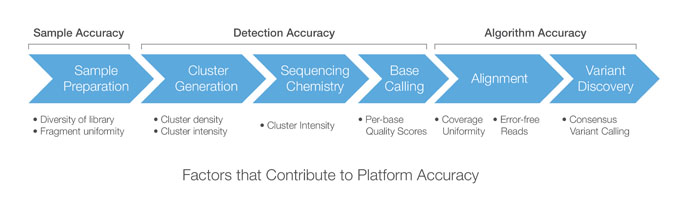
A typical sequencing workflow comprises sample/library preparation, cluster amplification, DNA sequencing, image analysis/base calling, read alignment, and variant discovery. If any of these steps generate poor results, the quality of the final data set is compromised.
With Illumina sequencing, each step in this process is optimized to deliver accurate data across a broad range of applications. Library preparation is an especially crucial step; using TruSeq or other high-quality Illumina library construction technologies can help ensure high sample quality and accuracy for any research project.
Platform Accuracy
Platform accuracy describes the overall accuracy of the sequencing workflow, accounting for each step of the process, from sample/library preparation through variant discovery. The sequencing workflow can be segmented into three main stages that each provide a unique accuracy contribution: Sample/Library Accuracy, Detection Accuracy, and Algorithm Accuracy.
Sample/Library Accuracy
Sample accuracy is associated with the library preparation step of the sequencing workflow. In this stage, DNA is fragmented for library construction. Each fragment in the library will eventually correspond to a sequencing read, so high fragment size uniformity and library diversity is important for achieving even coverage across the genome.
Errors that occur during library preparation, such as missing fragments due to a non-diverse library, cannot be identified by the sequencer. The portions of the genome not represented in the library will not be sequenced, leading to gaps in the data set.
In addition, quality scores do not reflect errors introduced during this step, as the sequencing signal will appear clean and error-free. The maximal achievable accuracy of most sequencing platforms is limited by the sample accuracy.
Thus it's critical to utilize high-quality library construction solutions such as TruSeq and other Illumina technologies.
Accurate Library Prep Technology
Our adapter ligation library prep technology, also known as TruSeq technology, has long been known for high coverage uniformity, precise strand information, and reliability. Now it's even better. Recent innovations have reduced the number of purification, sample transfer, and pipetting steps, improving workflow efficiency.
Learn More About Adapter Ligation
Illumina Sequencing Workflow
Learn more about the steps in the Illumina sequencing workflow: library preparation, sequencing, and data analysis. Discover how to plan your NGS experiment and view an example workflow.
Learn More
TruSeq Products
TruSeq kits provide simplicity, convenience, and trusted results for library preparation. Master-mixed reagents and a simple, automation-friendly workflow minimize hands-on time and reduce human error. Popular TruSeq library preparation products include:
Visit our library prep kit list page to find additional TruSeq products or explore other Illumina library preparation options.
TruSeq Library Prep Best Practices
Find recommended best practices for performing TruSeq library preparation and enrichment protocols.
A Tale of Two RNA Library Prep Kits
A critical comparison between two popular TruSeq RNA library prep kits reveals new information of interest to researchers conducting RNA sequencing studies.
Read InterviewCluster Density Tips
Optimal Cluster Density Best Practices
Watch Illumina scientists discuss how over- and underclustering can affect your sequencing data. Learn about common clustering issues and ways to prevent them.
View VideoHow to Achieve Consistent Cluster Density
This technical bulletin summarizes the resources and best practices to avoid underclustering and overclustering, and to achieve more consistent cluster densities.
Read Knowledge Article
Sequencing Systems
From benchtop to production-scale systems, find the right platform to advance your research.
Explore Our PlatformsDetection Accuracy
Detection accuracy accounts for the second stage of the sequencing workflow, comprising cluster generation, DNA sequencing, and primary data analysis. Any errors that occur during this stage are typically reflected in the quality scores.
Detection errors, unlike sample errors, can be tracked using the well-established per-base quality scores.
Detection errors can be improved by re-sequencing, single-read error correction, or encoding schemes.
Learn More About Quality ScoresAlgorithm Accuracy
Algorithm accuracy pertains to the secondary data analysis phase of the workflow, typically involving alignment and variant calling. The accuracy of the alignment method is critical.
Regardless of how high the quality of data is from the sequencing instrument, sub-optimal alignment will lead to a poor final data set, potentially with incorrectly placed mismatches, non-uniform coverage, and a high number of gaps. In turn, this can lead to high false positive and false negative rates. The variant calling method, by itself, also needs to be highly accurate for the same reasons.
Illumina offers user-friendly bioinformatics tools that enable researchers to perform accurate alignment and variant calling.
Explore Bioinformatics ToolsInterested in receiving newsletters, case studies, and information on new applications? Enter your email address below.
Featured Products
TruSeq Stranded Total RNA
A robust, highly scalable whole-transcriptome analysis (RNA-Seq) solution for a variety of species and sample types, including human, mouse, and formalin-fixed, paraffin-embedded (FFPE) tissue.
TruSeq DNA Nano
Generate whole-genome sequencing libraries and efficiently interrogate samples with limited available DNA.
TruSeq Small RNA Library Preparation Kits
These kits provide a simple, cost-effective solution for generating miRNA and small RNA sequencing libraries directly from total RNA, for any species.