Total RNA Sequencing
Introduction to Total RNA Sequencing
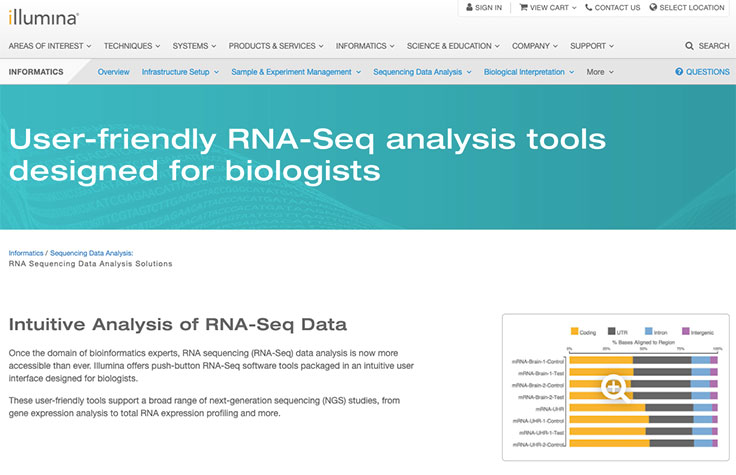
Whole-transcriptome analysis with total RNA sequencing (RNA-Seq) detects coding plus multiple forms of noncoding RNA. Total RNA-Seq can accurately measure gene and transcript abundance, and identify known and novel features of the transcriptome.
Total RNA-Seq provides optimal coverage in normal or low-quality samples. Species-specific ribosomal RNA probes can efficiently remove abundant RNA species. This leaves both fragmented and intact transcripts of interest for library capture.

Advantages of Total RNA Sequencing
Total RNA-Seq analyzes both coding and multiple forms of noncoding RNA for a comprehensive view of the transcriptome.
- Captures both known and novel features
- Allows researchers to identify biomarkers across the broadest range of transcripts
- Enables a more comprehensive understanding of phenotypes of interest
- Allows profiling of the whole transcriptome across a wide dynamic range

Transitioning from Whole-Genome Arrays to Total RNA-Seq
Dr. Avrum Spira, a pulmonary and critical care physician at Boston University Medical Center, started
a lab with the goal of analyzing samples with whole-genome gene expression arrays, to study
potential biomarkers for pulmonary disorders. Learn how Spira’s lab uses total RNA-Seq to uncover
novel and rare transcripts that may not be detectable on expression arrays.
Read Case Study.
Deciphering the Role of Long Non-Coding RNA in Cancer
Researchers at Biogazelle are using RNA-Seq to reveal how lncRNAs can be specific to cell types. They discovered that silencing certain lncRNAs can help destroy cancer cells.
Read Interview
Total RNA Sequencing Workflow
Illumina sequencing by synthesis (SBS) chemistry is the most widely adopted next-generation sequencing (NGS) technology, producing approximately 90% of global sequencing data.*
In addition to high-quality data, Illumina offers integrated whole-transcriptome sequencing workflows that simplify the entire process, from library preparation to data analysis and biological interpretation.

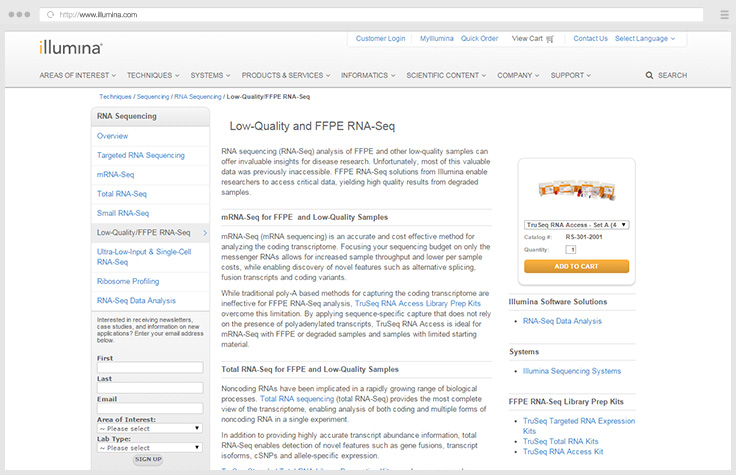
Library Prep for Total RNA-Seq
Our enhanced RNA-Seq library prep portfolio includes removal of abundant rRNA, so you can focus on high-value portions of the transcriptome.
Learn More
Using NGS to Study Rare Undiagnosed Genetic Disease
Whole-exome and transcriptome sequencing prove beneficial in uncovering mutations and pathways associated with rare disease
Learn More
Methods for RNA sequencing
This RNA sequencing methods guide provides Illumina solutions for profiling RNA, from targeted panels to the whole transcriptome. Illumina RNA sequencing workflows seamlessly integrate library prep, sequencing, and data analysis to support transcriptome research.
Download guideFeatured Products
Related solutions
Cancer research: gene expression studies
Monitoring gene expression changes in both coding and noncoding RNA biomarkers with total RNA-Seq can help researchers understand which variants affect tumor classification and progression. Learn more about cancer RNA-Seq.
Agrigenomics: deep transcriptome sequencing
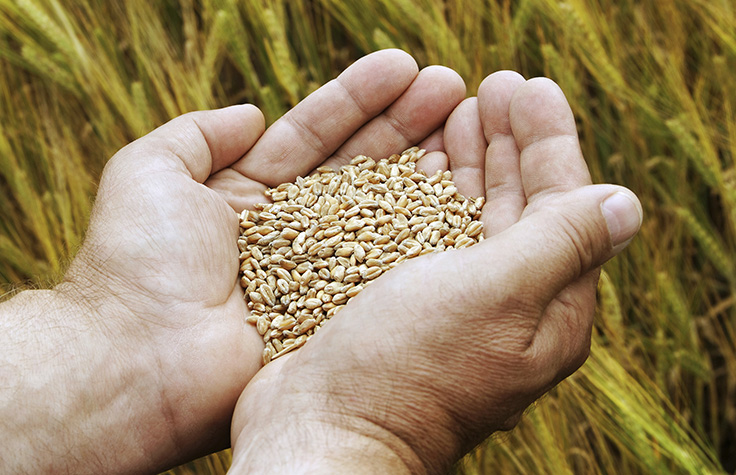

Plants have large, repetitive genomes, which can make it difficult to sequence weakly expressed genes. Ultradeep RNA sequencing can facilitate analysis of these genes. Learn more about plant sequencing.
Complex disease biomarker identification

Total RNA-Seq allows complex disease researchers to study coding and multiple forms of noncoding RNA in a single analysis, providing visibility to a broad range of potential disease-associated biomarkers. Learn more about complex disease research.
Spatial transcriptomics
Spatially resolve transcriptional activity in complex tissue architectures using RNA-Seq. Learn how spatial transcriptomics combines molecular profiling with spatial context.