Microarray Technology
Bead-Based Microarray Technology
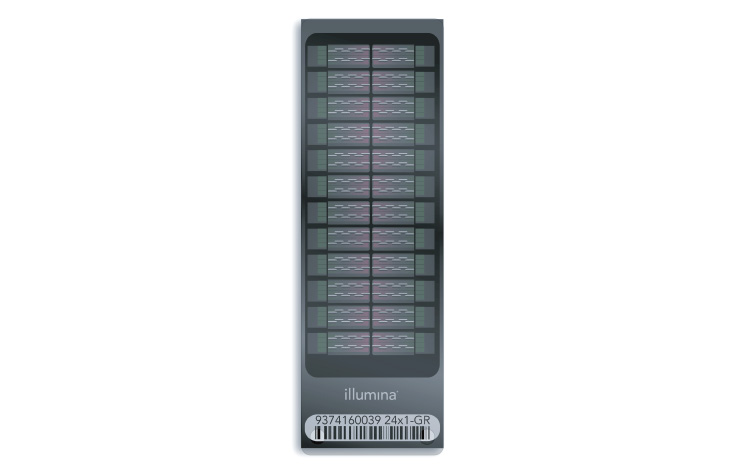
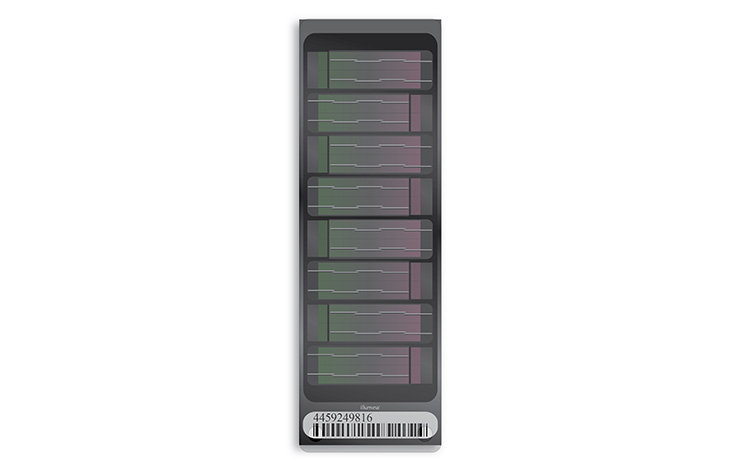
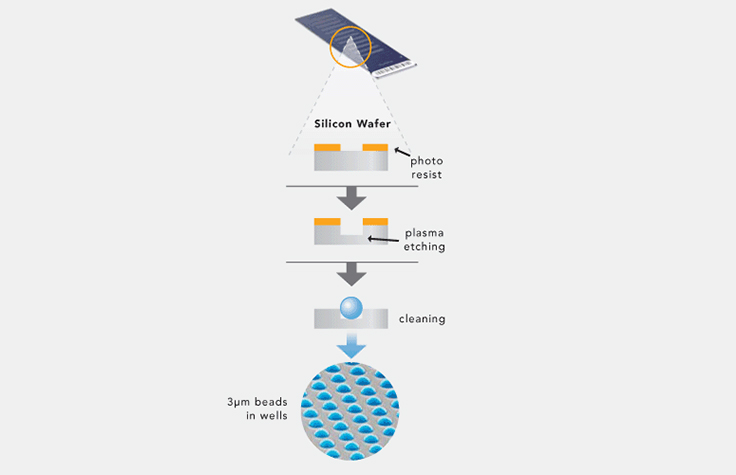
Illumina microarray technology (also known as BeadArray technology) uses silica microbeads. On the surface of each array, or BeadChip, hundreds of thousands to millions of genotypes for a single individual can be assayed at once. These tiny silica beads are housed in carefully etched microwells and coated with multiple copies of an oligonucleotide probe targeting a specific locus in the genome.
How Do Illumina Microarrays Work?
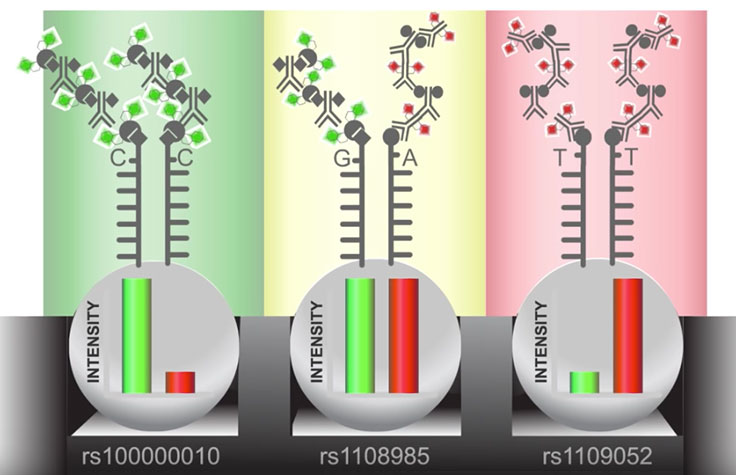
As DNA fragments pass over the BeadChip, each probe binds to a complementary sequence in the sample DNA, stopping one base before the locus of interest. Allele specificity is conferred by a single base extension that incorporates one of four labeled nucleotides. When excited by a laser, the nucleotide label emits a signal. The intensity of that signal conveys information about the allelic ratio at that locus.
Trusted Microarray Technology
Trusted data quality and exceptional coverage of valuable genomic regions make our Infinium genotyping arrays the assay of choice by leading institutions for high- throughput research screening and large-scale research programs. Infinium chemistry produces exceptional data quality and call rates, as well as consistent reproducibility. We offer a broad range of microarray options based on SNP complexity and sample throughput.
Infinium-Powered Array Progress
Since the launch of our first BeadChip, we have innovated microarray solutions to help researchers advance science.
View Video
Methylation Array Technology
Infinium methylation technology allows highly accurate and precise quantification of methylation levels in the genome. Use the Infinium methylation assay to detect cytosine methylation at the level of single CpG sites.
Multiomics multiplies your discovery power
See how you can use multiomics to better connect genotype to phenotype and obtain a full cellular readout not found through single omics approaches.

Popular Products
How scientists are using microarray technology
Read how scientists are using arrays to further their genomic research across a diverse field of applications.
Exploring variants of unknown significance
Read how Dr Benjamin Darbro at the University of Iowa Carver College of Medicine, uses a wide range of cytogenetic tools, including microarrays and more, to explore gene networks involved in neurodevelopmental disorders.
Identification of selection signatures and genetic diversity in the sheep
In this research paper, scientists make significant contributions to a better understanding of the genetic structure and production potential of Eşme sheep to optimize and inform breeding programs.
How the methylome relates to clinical research outcome
In this spotlight, Dr Kathleen Barnes talks about how genetic factors can influence disease in populations as well as factors influencing its global distribution using methylation arrays.
iScan System Global Screening Array Research Bundle
Microarray Selection Tools
Library Prep & Array Kit Selector
Find the right microarray or library prep kit for your needs. Filter by method, species, and more. Compare, share, and order kits.
Featured microarray solutions
Learn about the Infinium array product line that provide solutions for unparalleled genomic access and accuracy to detect genetic and epigenetic variation.
Featured webinars
A robust, high-throughput genotyping platform
See how scientists developed a new multi-species platform based on the Illumina Infinium II array chemistry targeting approximately 3000 SNPs in wheat, oat and barley.
Mouse DNA methylation arrays
Watch how a mouse DNA methylation array can accelerate high sample-throughput studies in this important model organism.
Microarray Technology Tips
DNA Strand Designations
Learn about the different strand designations found in Illumina array manifests and GenomeStudio projects.
Guidelines for Identifying TOP/BOT Strand and A/B Allele
A step-by-step method to help you understand this nomenclature for single nucleotide polymorphisms.
How to Interpret DNA Strand and Allele Information for Array Data
When comparing genotyping data, it is important to use the same DNA strand designation.
Related applications
Microarray techniques
SIllumina microarrays offer high-quality data and exceptional genomic coverage to propel genomic studies of any size.
Human genotyping research
Human genotyping arrays are ideal for processing thousands of samples to identify variants associated with traits and disease.
Methylation array analysis
Methylation arrays enable high-throughput, quantitative interrogation of methylation sites across the genome at single-nucleotide resolution.
Microarray data analysis
Software tools for array experimental design, sample tracking, and analysis of microarray data.
Featured publications
Identifying genetic variants in metabolic dysfunction
In this research paper, scientists investigate genes and variants underlying metabolic dysfunction-associated fatty liver diseases in a Korean adult population at a genome-wide level.
Prenatal microarray in high-risk pregnancies
Scientists show that chromosomal microarray analysis can enhance the prenatal diagnostic accuracy by detecting submicroscopic copy number variants not visible with conventional methods.
SNP array profiling for breast cancer
Read how researchers used SNP array profiling to gain a better understanding of breast cancer etiology to facilitate predictive therapies and improve prognostic assessments.