QTL Analysis and Mapping

What is QTL Analysis?
A quantitative trait locus (QTL) is a region of DNA associated with a specific phenotype or trait that varies within a population. Typically, QTLs are associated with traits with continuous variance, such as height or skin color, rather than traits with discrete variance, such as hair or eye color.
QTL mapping is a statistical analysis to identify which molecular markers lead to a quantitative change of a particular trait. Since a single locus may include many variants, imputation or whole-genome sequencing is a key prerequisite for QTL mapping to enable precise identification of the contributing molecular marker. QTLs have been expanded to include variants that act at different levels throughout the genotype-to-phenotype continuum.
Types of QTL Analysis
QTL analysis is an effective means of annotating variants that are associated with disease. By understanding the functional effects of variants, it allows for the distinction between variants that are involved with disease, from those that are correlated with disease. By leveraging different QTL analyses, the network of molecular interactions of variants and the genes they affect begin to come into view, and provide evidence for which underlying genes and pathways are truly driving disease. This enables the investment of time, resources, and funding in targets that are most likely to be involved with disease.
eQTL
Expression quantitative trait loci (eQTL) are genetic variants that affect the expression of one or more genes. An eQTL can act in cis (locally) or in trans (at a distance, eg, on a different chromosome) to its gene target. eQTL mapping requires genome-wide genotyping by arrays or WGS and gene expression analysis by RNA-Seq for each sample.
Learn more about:
meQTL
Methylation quantitative trait loci (meQTL) are genetic variants that affect patterns of DNA methylation. meQTLs can influence methylation across extended genomic regions. meQTL mapping requires genome-wide genotyping and DNA methylation analysis with either arrays or sequencing.
Learn more about:
caQTL
Chromatin accessibility quantitative trait loci (caQTL) are genetic variants that affect nucleosome packing, positioning, and chromatin accessibility. caQTL mapping requires genome-wide genotyping by arrays or WGS and chromatin accessibility analysis by a method such as ATAC-Seq or Hi-C.
Learn more about:
bQTL
Binding quantitative trait loci (bQTL) are genetic variants that affect transcription factor binding. bQTL mapping requires genome-wide genotyping by arrays or WGS and ChIP-Seq.
Learn more about:
pQTL
Protein quantitative trait loci (pQTL) are genetic variants that affect the quantity of that particular protein. pQTL requires genome-wide association studies (GWAS) using arrays, whole-genome (WGS), or whole-exome sequencing (WES).
Learn more about:
QTL Analysis Applications
We support the following workflows for in-depth analysis of basic and clinical research data:
Complex Disease
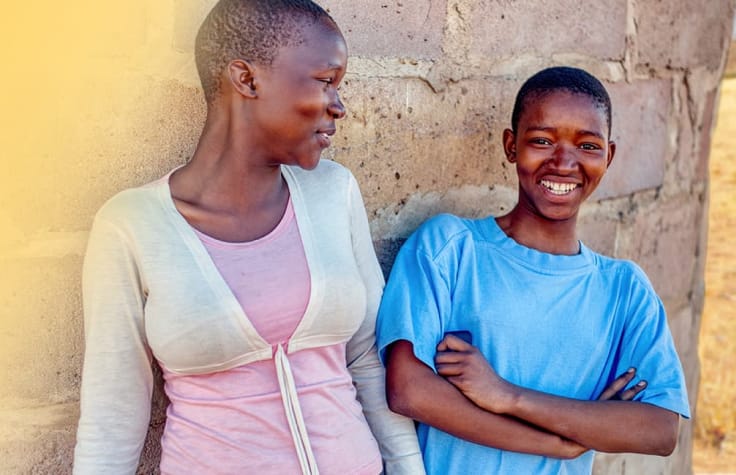

From variant discovery to risk profiling, modern genomics tools are helping researchers investigate the molecular etiology of complex diseases.
Genetic & Rare Diseases
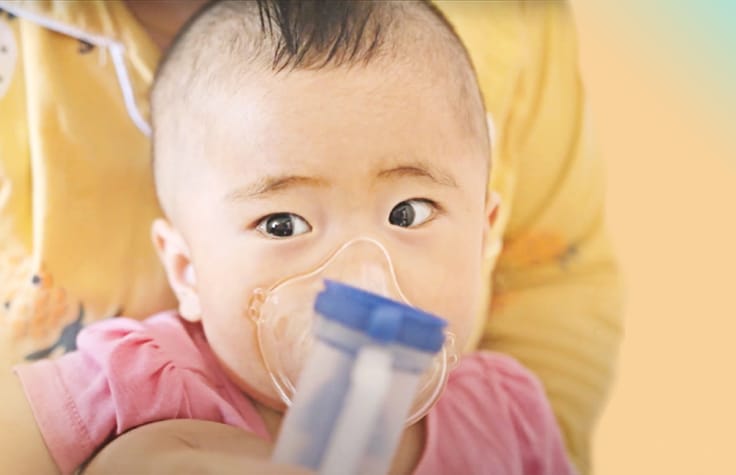

NGS technology is helping to drive breakthroughs in genetic disease testing by facilitating early detection and diagnosis.
Agrigenomics
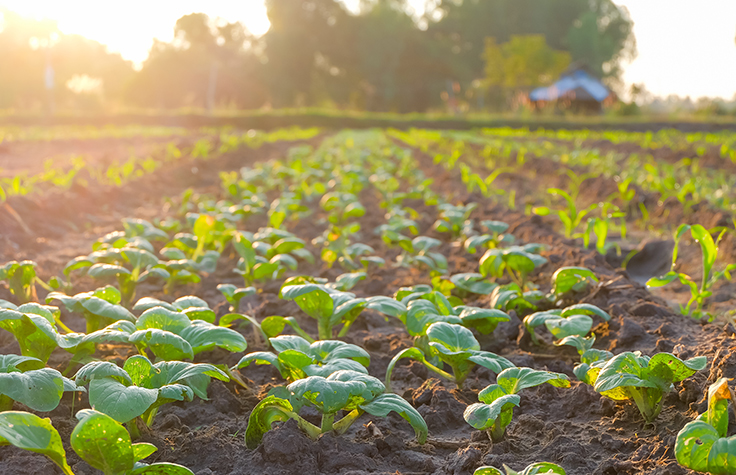

Genomics is helping farmers and breeders develop more productive crops and livestock, improve sustainability, and meet the growing challenges of feeding the world.
Cancer Research
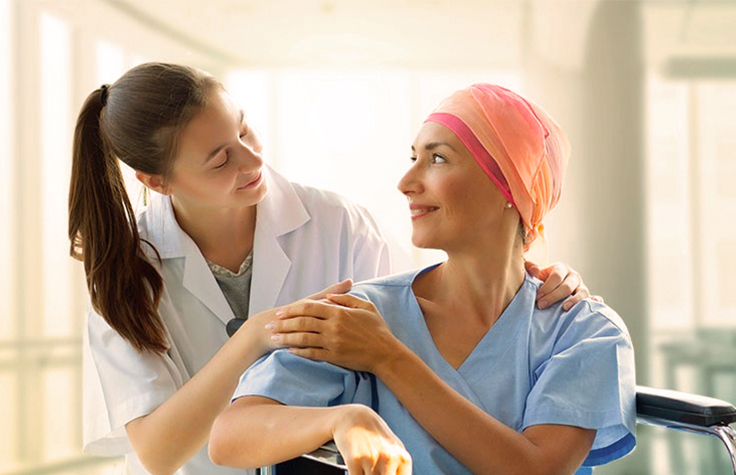

Today’s genomic technologies can help you uncover new insights into cancer biology and etiology for tomorrow’s research breakthroughs.
QTL Analysis FAQ
Webinar: From Variants to Function with Multiomics
In this session, hear from three panelists who will share how we can begin to understand what variants mean physiologically through a multiomics approach. You'll learn how powerful combinations of high-throughput experimental assays, single-cell approaches, and computational analyses are accelerating the ability to link variants to function, and by extension, link genotype to phenotype.
Watch Webinar
Related Content
Gene Target Identification
NGS-based methods reveal a variant's effects on gene expression to better characterize disease mechanisms.
Technical Note: QTL Analysis Software Tools for Illumina Data
Illumina provides data analysis software that supports data integration, such as genotyping with gene expression for eQTL analysis.
GenomeStudio Software
Visualize and analyze data generated on Illumina array platforms with GenomeStudio Software. This powerful solution supports the genotyping analysis of microarray data.
Video: Multiomics is Revolutionizing Research
Learn why these research leaders decided to use multiomics in their work.
Whole-Genome Sequencing
Whole-genome sequencing delivers a comprehensive view, ideal for discovery applications. Newer genome sequencers perform WGS more rapidly than ever.
COVID-19 Immune Profiling
See how to unravel the complexities of the disease and reveal future therapeutic targets.