Amplicon Sequencing
Introduction to Amplicon Sequencing
Amplicon sequencing is a highly targeted approach that enables researchers to analyze genetic variation in specific genomic regions. The ultra-deep sequencing of PCR products (amplicons) allows efficient variant identification and characterization. This method uses oligonucleotide probes designed to target and capture regions of interest, followed by next-generation sequencing (NGS).
Flexible, Accessible Sequencing Instruments
Benchtop sequencing systems enable novice users to perform targeted sequencing across a range of applications.
Advantages of Amplicon Sequencing
Amplicon sequencing is useful for the discovery of rare somatic mutations in complex samples (such as tumors mixed with germline DNA). Another common application is sequencing the bacterial 16S rRNA gene across multiple species, a widely used method for phylogeny and taxonomy studies, particularly in diverse metagenomics samples.
- Enables researchers to efficiently discover, validate, and screen genetic variants using a highly targeted approach
- Supports multiplexing of hundreds to thousands of amplicons per reaction to achieve high coverage
- Delivers highly targeted resequencing even in difficult-to-sequence areas, such as GC-rich regions
- Allows flexibility for a wide range of experimental designs
- Reduces sequencing costs and turnaround time compared to broader approaches such as whole-genome sequencing
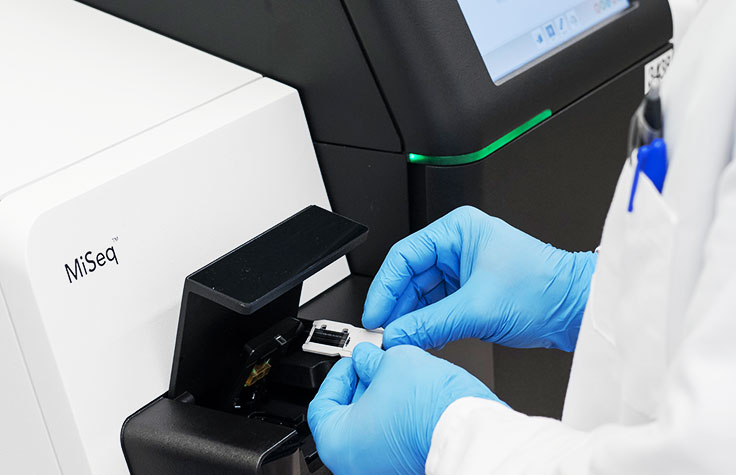

Highly Targeted, Multiplexed Analysis
Amplicon sequencing allows researchers to sequence targets ranging from a few to hundreds of genes in a single run. This ultra-high multiplexed PCR approach expedites research by assessing multiple genes simultaneously. Libraries can be prepared in as little as 5–7.5 hours and sequenced in 17–32 hours.
Amplicon sequencing enables a wide range of research applications for the discovery, validation, or screening of genetic variants.
Recommended Workflow for Amplicon Sequencing
Content Selection and Library Prep
DesignStudio Custom Assay Designer
Web-based custom assay design tool.
AmpliSeq for Illumina Custom DNA Panel
Targeted custom research panels optimized for specific targets or genomic content of interest.
Sequencing
MiSeq i100 Series
Our fastest, simplest Illumina benchtop system for amplicon sequencing.
Data Analysis
BaseSpace Sequence Hub (DNA Amplicon App) and BaseSpace Variant Interpreter
Amplicon analysis and variant detection tools.

Methods for microbial single-genome sequencing
This guide offers library preparation, sequencing systems, and corresponding data analysis apps for NGS workflows that enable discoveries and provide insights to understand your microbe of interest better. These workflows can enable the full array of microbiology and infectious disease research applications.
Download guideComprehensive Amplicon Sequencing Workflow
As the global leader in NGS technology, Illumina has installed over 25,000 instruments worldwide and its technology is referenced in more than 400,000 peer-reviewed publications—five times more than all other NGS providers combined.*
Illumina offers integrated amplicon sequencing workflows that simplify the entire process, from library preparation to data analysis and biological interpretation.
Click on the below to view products for each workflow step.
Delivers a range of ready-to-use and custom panels for simple, flexible targeted resequencing that provides high-quality data you can trust.
Illumina DNA PrepA fast, integrated workflow for a variety of applications, from human whole-genome sequencing to amplicons, plasmids, and microbes.
TruSight Tumor 15Focused sequencing research panel to assess 15 genes commonly mutated in solid tumors in a single assay, with a simple, rapid workflow.
Nextera XT DNA Library Prep KitLibrary preparation for small genomes (bacteria, archaea, viruses), amplicons, and plasmids in less than 90 minutes.
Illumina targeted sequencing panels facilitate clinical cancer research by providing expert-defined gene content plus simple data analysis and reporting options.
Library Prep Kit SelectorDetermine the best kit for your needs based on project type, starting material, and method or application.
DesignStudio Assay Design ToolAn easy-to-use online software tool that provides dynamic feedback to optimize probe designs.
Affordable sequencing power for targeted or small genome sequencing in any lab.
MiniSeq SystemSupports a broad range of targeted DNA and RNA applications for examining single genes or entire pathways.
MiSeq i100 SeriesAccessible benchtop systems with remarkably simple operations delivering same-day, accurate results.
NextSeq 550 SystemProvides the flexible power you need for whole-genome, transcriptome, and targeted resequencing.
Compare sequencing platforms and identify the best system for your lab and applications.
Sequencing ReagentsFind kits that include sequencing reagents, flow cells, and/or buffers tailored to each Illumina sequencing system.
The Illumina genomics computing environment for NGS data analysis and management.
Local Run ManagerAn on-premises software solution for creating sequencing runs, monitoring run status, and analyzing data.
DNA Amplicon AppStreamlined analysis of NGS data enriched for particular target sequences using amplicon reads.
16S Metagenomics AppAnalyzes DNA from amplicon sequencing of prokaryotic 16S small subunit rRNA genes and provides visuals of taxonomic classification.
A genome browser developed by the Broad Institute of MIT and Harvard that displays NGS data for complex variant analysis.
BaseSpace Variant InterpreterLeverages leading annotation databases and a powerful filtering interface, enabling researchers to rapidly identify disease-associated variants.
BaseSpace Correlation EngineA growing library of curated genomic data to support researchers in identifying disease mechanisms, drug targets, and biomarkers.
Speed and simplicity with the MiSeq i100 Series benchtop sequencers
The MiSeq i100 Series delivers our fastest run times yet, breakthrough simplicity, and supported sample-to-analysis workflows for amplicon sequencing.
View System
Related Solutions
Cancer Gene Sequencing
Targeted sequencing enables researchers to focus on select genes or amplicons that have known associations with cancer. Both predesigned and custom panels are available. Learn more about targeted cancer sequencing.
Genetic and Rare Diseases
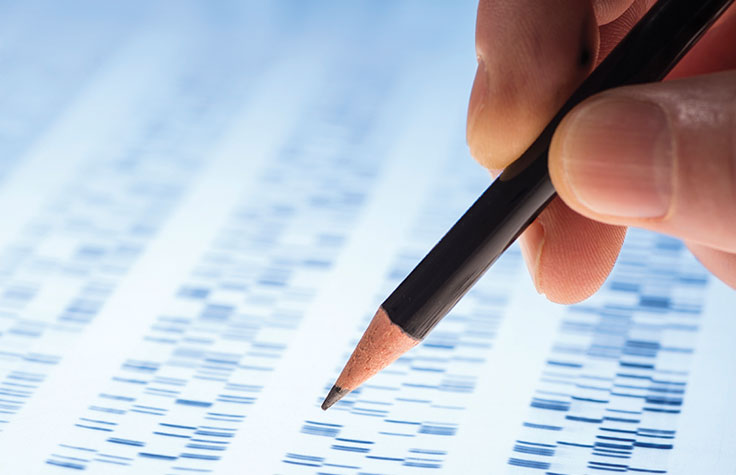

NGS technology can help rapidly identify causative variants associated with rare and inherited genetic disorders. In contrast, traditional methods are often costly and require extensive testing. Learn more about NGS for genetic diseases.
16S rRNA Sequencing
16S ribosomal RNA (rRNA) sequencing is a common amplicon sequencing method used to identify and compare bacteria within a given sample. It can identify strains that might not be found using other methods. Learn more about 16S rRNA sequencing.
Plant and Animal Sequencing
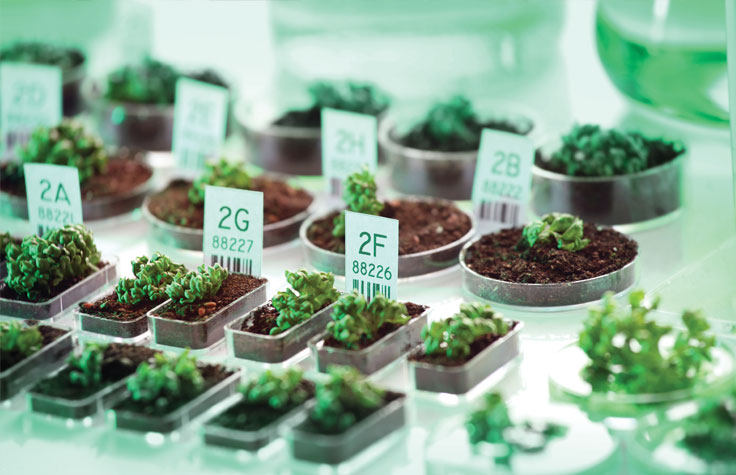

Targeted resequencing can uncover genetic variation in animals and plants. These variants may represent beneficial mutations that can help inform breeding decisions, or mutations linked to disease susceptibility. Learn more about plant and animal sequencing.
Interested in receiving newsletters, case studies, and information on genomic analysis techniques? Enter your email address.
Additional Resources
Demonstrated 16S rRNA Sequencing Protocol
Illumina offers a demonstrated amplicon-based library prep protocol for 16S metagenomic sequencing.

AmpliSeq for Illumina on the iSeq 100 System
Find step-by-step guidance designed to help you transition your workflows to AmpliSeq for Illumina panels on the iSeq 100 System.

Transitioning from TruSeq Custom Amplicon
Our detailed guide is designed to help you transfer projects from TruSeq Custom Amplicon to AmpliSeq for Illumina panels.
*Data calculations on file. Illumina, Inc., 2022.